Hello GENI
Updated 3/1/2021
Overview:Welcome to GENI. This page will guide you through your first GENI experiment.This is a very simple tutorial with one server and one client that are connected with a Layer 2 link. 
|
|
Prerequisites:For this tutorial you need a GENI Experimenter Portal account and be a member of at least one project.
|
Tools:All the tools will already be installed at your nodes. For your reference we are going to use:
|
|
Where to get help:For any questions or problem with the tutorial please email geni-users@googlegroups.com |
Step-by-step Instructions
 |
Step 1: Get Ready:
The first thing we need to do is login to the portal.
Tip: Start typing the name of your institution and see the list become smaller. |
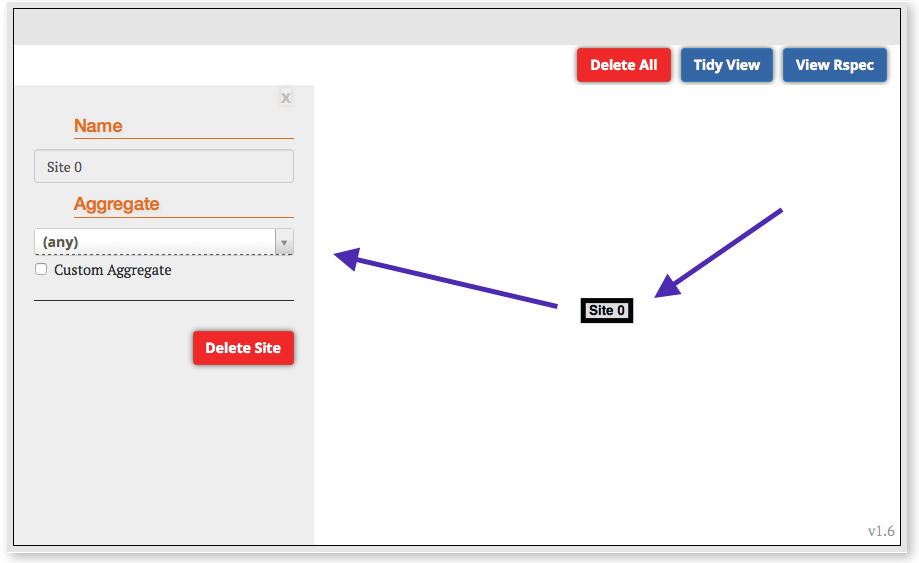
Step 2: Launch your experiment:
|
|
 |
Step 3: View Results:
For this example experiment we used the install script facility to automatically install the necessary software and kick-off the experiment. In this very simple setup, we have installed and launched a web server as well as an iperf server, on the server host. On the client, we have started some processes to test both of these services. To view the results of this experiment:
|
Optional Step 4: Manually generate traffic:
While conducting experiments in GENI, you will often want to run commands directly on the nodes. In this optional step, you will log in to a node and issue commands directly to it.- Follow these instructions and log in to the client node
- When you have successfully logged in, run this command:
iperf -c server -P 2
This task shouldn't take more than 30 seconds. Change the number after the ` -P ` argument and watch how the performance is affected while you change the number of parallel TCP connections. - Scroll all the way down the server iperf log, and look at the logs for your transfers
 |
Step 5: Cleanup experiment:
After you are done with your experiment, you should always release your resources so that other experimenters can use the resources. In order to cleanup your slice :- Press the Delete button in the bottom of your Jacks canvas.
What's next?
Congratulations! You have finished your first GENI Experiment. Now that you are more familiar with GENI concepts you can:
- continue with more advanced tutorials
- learn more about how to use Jacks by following the A First Experiment Using GENI and Jacks Tool
- start with your own experiment. This tutorial showed you the basic steps for running an experiment. Use Jacks to create your own topology, take a look at the instructions about how to write your own install scripts to automate your experiment.
Last modified 3 years ago
Last modified on 03/03/21 11:52:52
