| Version 5 (modified by , 11 years ago) (diff) |
|---|
GIMI & GEMINI on one slice
InstaGENI:
Creation:
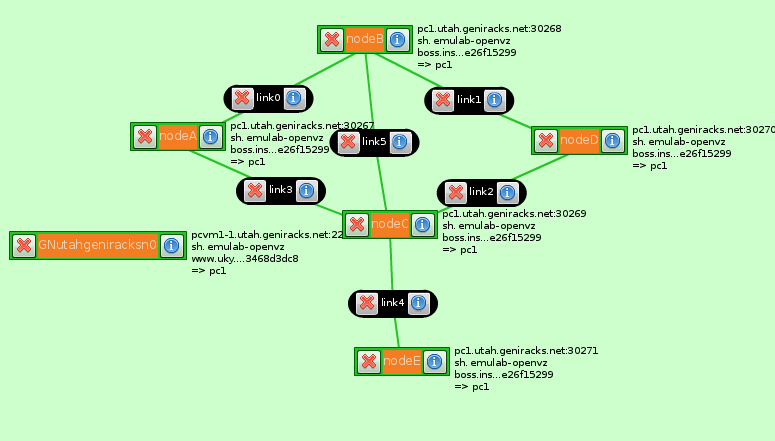
I started with this rspec which was created to use GIMI on InstaGENI. I created a couple slices & used that rspec in Flack with an InstaGENI aggregate. I then added GEMINI extensions through Flack. Here is an example of the final rspec at Utah InstaGENI. Below is the topology created.
GEMINI:
I was able to initialize and instrumentize slices via the GENI Desktop and gdesktop scripts. These slices were able to use all the GENI Desktop features.
GIMI:
In Labwiki I ran the four experiment template scripts on the slices. I was even able to see changes on the graphs shown by the GENI Desktop while Labwiki was running experiments.

ExoGENI:
Creation:
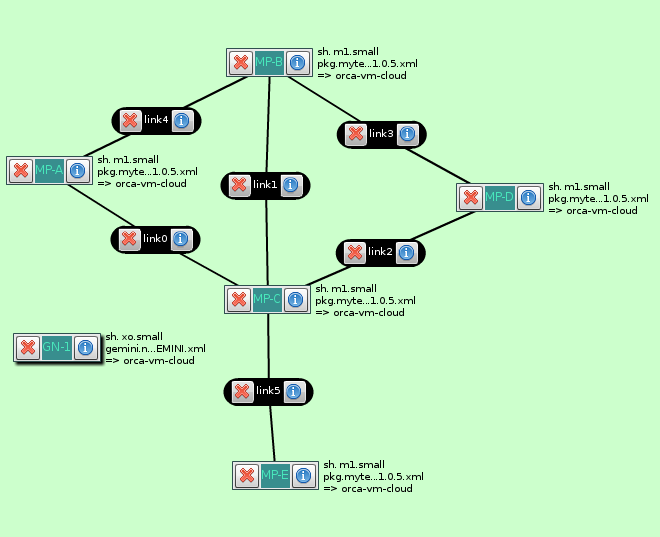
I used this rspec to create ExoGENI slices in Flack that had both GIMI and GEMINI. Below is the topology created.
GEMINI:
Since the stable version of the GENI Desktop does not currently handle ExoGENI slices, I used the development version of the GENI Desktop and the gdesktop scripts to initialize & instrumentize the slices. Within the GENI Desktop I could not see graphs or tables. I was able to SSH into the nodes via the GENI Desktop.
GIMI:
In Labwiki I was able to run the four experiment template scripts on the slices.
Attachments (36)
- GnGInstTopology.png (49.1 KB) - added by 11 years ago.
-
GIMI5nodeInstaGENInoAM.xml (7.8 KB) - added by 11 years ago.
GIMI with InstaGENI rspec
-
GnGInstaRspec.xml (11.8 KB) - added by 11 years ago.
Utah InstaGENI rspec with GIMI & GEMINI
- GnGSliceOnGENIDesktop.png (311.1 KB) - added by 11 years ago.
- LabwikiAndGENIDesktop.png (365.7 KB) - added by 11 years ago.
-
GIMI_GEC17wGN.xml (6.2 KB) - added by 11 years ago.
GIMI ExoGENI with GEMINI
- GnGExoTopology.png (44.6 KB) - added by 11 years ago.
- GENIDesktopLogin.png (30.9 KB) - added by 11 years ago.
- GENIDesktopLogin2.png (47.7 KB) - added by 11 years ago.
- GENIDesktopLoginSSHPass.png (51.4 KB) - added by 11 years ago.
- GENIDesktopClickInstrument.png (33.9 KB) - added by 11 years ago.
- GENIDesktopInstrumentized.png (37.5 KB) - added by 11 years ago.
- GENIDesktopSSHWindow.png (24.4 KB) - added by 11 years ago.
- GENIDeskopSSH.png (35.2 KB) - added by 11 years ago.
- GENIDesktopUntrustedSSH.png (64.0 KB) - added by 11 years ago.
- GENIDesktopLaunchGraphs.png (23.3 KB) - added by 11 years ago.
- GENIDesktopGraph.png (201.1 KB) - added by 11 years ago.
- GENIDesktopLaunchGN.png (106.5 KB) - added by 11 years ago.
- GENIDesktopActiveMeasure.png (197.4 KB) - added by 11 years ago.
- LabWikiSendInfo.png (24.8 KB) - added by 11 years ago.
- LabwikiTypeStep.png (17.9 KB) - added by 11 years ago.
- LabWikiStep1Save.png (44.2 KB) - added by 11 years ago.
- LabWikiStep1Drag.png (58.2 KB) - added by 11 years ago.
- GENIDesktopPickSlice.png (38.7 KB) - added by 11 years ago.
- LabWiki.png (75.3 KB) - added by 11 years ago.
- GENIDesktopInitialize.png (14.6 KB) - added by 11 years ago.
- LabWikiandGEMINIGraphs.png (408.6 KB) - added by 11 years ago.
- LabWikiEnd.png (56.9 KB) - added by 11 years ago.
- LabWikiExpRunning.png (121.4 KB) - added by 11 years ago.
- LabWikiStep1Input.png (74.1 KB) - added by 11 years ago.
- LabWikiStart2.png (54.9 KB) - added by 11 years ago.
- gdesktopInit.png (50.1 KB) - added by 11 years ago.
- gdesktopInstrument.png (52.6 KB) - added by 11 years ago.
- GENIDesktopStable.png (45.2 KB) - added by 11 years ago.
- GENIDesktopDevelopment.png (59.9 KB) - added by 11 years ago.
- GnGonExo.png (278.6 KB) - added by 11 years ago.