| Version 16 (modified by , 12 years ago) (diff) |
|---|
E. Push to iRODS

D.1 Overview
In GIMI, iRODS is used as the repository for measurement data. At the moment our iRODS data system consist of three servers (RENCI, NICTA, and UMass) and a metadata catalog (located at RENCI).
D.2 Storing data on iRODS
- After successful completion of an experiment the measurement data from the experiment have been stored in iRODS. We will now look into two options on how this data can be handled.
D.2.1 iRODS command line tools in the user work space
- The GENI User Workspace comes already with the iRODS command line tools installed.
- If you have not done so far use iinit to create a file that contains your iRODS password in a scrambled form. This will then be used automatically by the icommands for authentication with the server.
$ iinit
You will be prompted for a password. Please enter the one given on the paper handout.
- The iRODS client uses a configuration file (~/.irods/.irodsEnv) that sets certain parameter for the icommands. Here is and example:
# iRODS personal configuration file. # # This file was automatically created during iRODS installation. # Created Thu Feb 16 14:06:27 2012 # # iRODS server host name: irodsHost 'emmy9.casa.umass.edu' # iRODS server port number: irodsPort 1247 # Default storage resource name: irodsDefResource 'iRODSUmass2' # Home directory in iRODS: irodsHome '/geniRenci/home/gimiXX' # Current directory in iRODS: irodsCwd '/geniRenci/home/gimiXX' # Account name: irodsUserName 'gimiXX' # Zone: irodsZone 'geniRenci'
- Retrieve file from your iRODS home directory into user workspace.
$ iget <file_name>
Assuming you wanted to retrieve a file called gimitesting.sq3 the command would look as follows:
$ iget gimitesting.sq3
- Store data from user workspace into your iRODS home directory.
$ iput <file_name>
- Add metadata information to stored file.
$ imeta add -d <file_name> <name> <value> <unit>
For example if you wanted to add the slice name of the experiment as metadata for file gimi01-ping_all.sq3, the command would look as follows:$ imeta add -d gimi01-ping_all.sq3 SliceName gimi01
(<unit> is not required.)
D.2.2 iRODS web interface
iRODS also provides a nice and easy to use web interface, which we will explore in the following.
- Point the browser in your user workspace to the following link: https://www.irods.org/web/index.php
- Input the following information to sign in:
Host/IP: emmy9.casa.umass.edu Port: 1247 Username: as given on printout Password: as given on printout
- The following screenshot shows an example for the web interface:
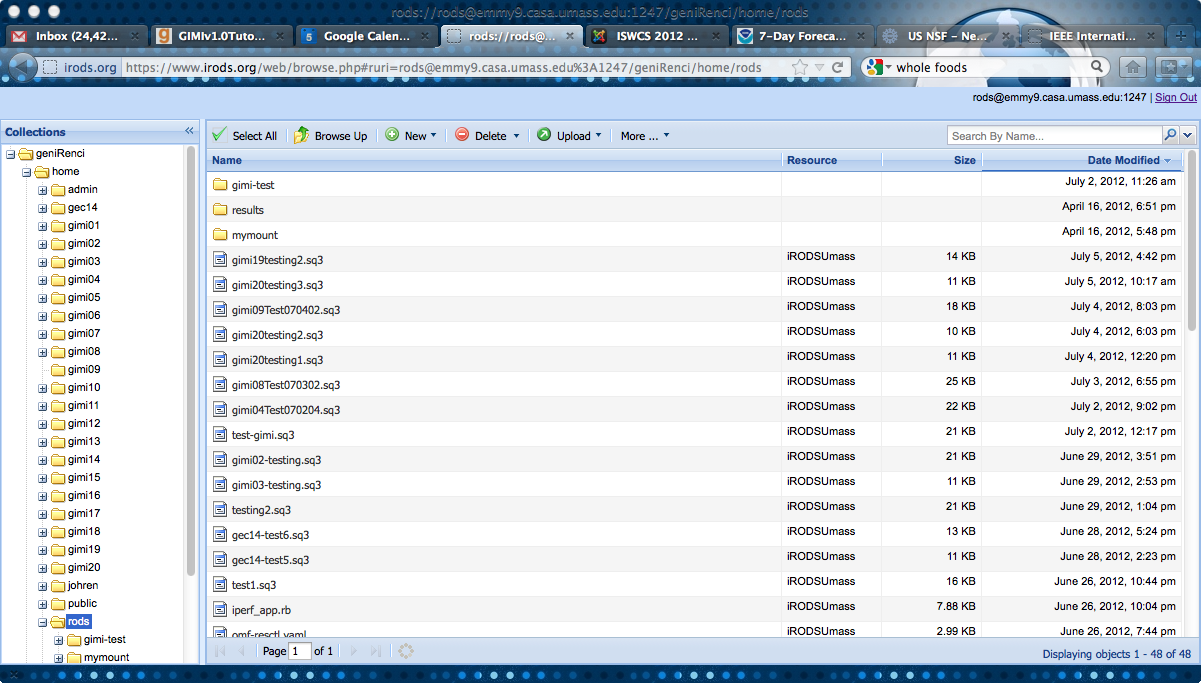
D.3 IRODS and OML
In GIMI we have enabled an option in OML2.8 that allows the execution of script. We use this functionality to automatically save measurement results in IRODS after a measurement has successfully completed.
D.4 DISCLAIMER
The iRODS service we are offering within the scope of GIMI does NOT guarantee 100% reliable data storage (i.e., we do NOT back up the data). If you are performing your own experiments and want to use iRODS you are absolutely welcome but be aware that we do NOT guarantee recovery from data loss.
6) Move selected collected measurements and other artifacts from storage service to long-term archive service
- identify archived objects with peristent identifier
- include policy for sharing with others
- allow retrieval for further analysis and visualization
http://groups.geni.net/geni/wiki/GIMIv1.1Tutorial/Observe Back to previous step
http://groups.geni.net/geni/wiki/GIMIv1.1Tutorial/Analyze Forward to next step
http://groups.geni.net/geni/wiki/GIMIv1.1Tutorial/ Back to tutorial main page
Attachments (1)
-
GIMI_workflow_E.jpg (39.5 KB) - added by 12 years ago.
Workflow E
Download all attachments as: .zip
