| Version 19 (modified by , 12 years ago) (diff) |
|---|
GENI Racks and Meso-scale OpenFlow Interoperability
This page describes an interoperability experiment that has been completed between Meso-scale OpenFlow sites, ExoGENI and InstaGENI OpenFlow Resources. This experiment used the OpenFlow VLAN 1750 was used at each Meso-scale site and the GENI Core OpenFlow VLAN 3716. In this experiment the following resources are reserved:
ExoGENI:
- Two BBN ExoGENI rack VMs on OpenFlow shared VLAN 1750
- Two RENCI ExoGENI rack VMs on OpenFlow shared VLAN 1750
- Two BBN Campus VMs that have access to OF VLAN 1750 via flowspaces through the ExoGENI rack OF switch. (These OF flowspaces are managed with the ExoGENI FOAM aggregate).
InstaGENI:
- Two InstaGENI rack VMs on OpenFlow shared VLAN 1750
- Utah PG Campus host that has access to OF VLAN 1750 via flowspaces through the InstaGENI rack OF switch. (These OF flowspaces are managed with the InstaGENI FOAM aggregate.)
Meso-scale:
- Three meso-scale OpenFlow Aggregates: Rutgers, BBN and Clemson.
- Three meso-scale compute resources sites: 1 WAPG at Rutgers(Internet2), 1 MyPLC at BBN, and 1 MyPLC at Clemson (NLR).
- NRL and Internet2 OpenFlow VLAN 3716 is used for all hosts in experiment1 to exchange traffic
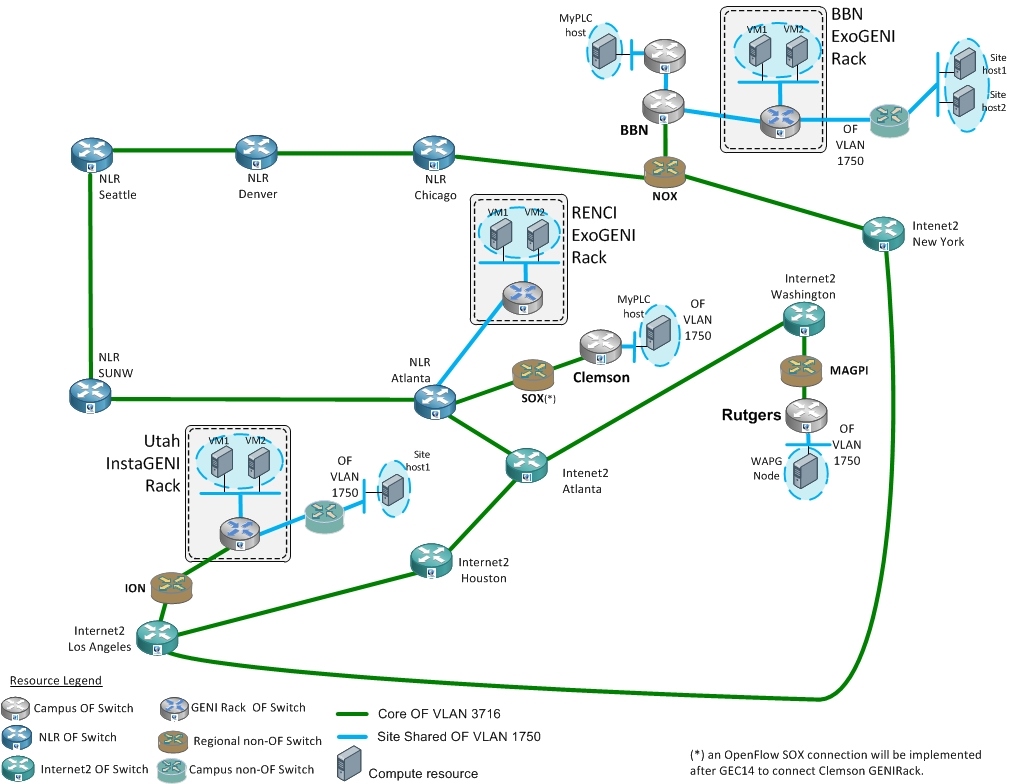
The following diagram describes the Meso-scale and GENI Racks interoperability experiment captured in this page:
Omni configuration settings
The Omni configuration defined the pgeni.gpolab.bbn.com to be used for the credentials and as a slice authority. To simplify the experiment setup, the following aggregate manager nick_names were defined in the omni_config:
#ExoGENI Compute and OF Aggregates Managers exobbn=,https://bbn-hn.exogeni.net:11443/orca/xmlrpc exorci=,https://rci-hn.exogeni.net:11443/orca/xmlrpc exosm=,https://geni.renci.org:11443/orca/xmlrpc of-exobbn=,https://bbn-hn.exogeni.net:3626/foam/gapi/1 of-exorci=,https://rci-hn.exogeni.net:3626/foam/gapi/1 #InstaGENI Compute and OF Aggregates Managers ig=,http://utah.geniracks.net/protogeni/xmlrpc/am of-ig=,https://foam.utah.geniracks.net:3626/foam/gapi/1 pg=,http://www.emulab.net/protogeni/xmlrpc/am #Meso-scale Compute and OF Aggregates Managers of-bbn=,https://foam.gpolab.bbn.com:3626/foam/gapi/1 of-clemson=,https://foam.clemson.edu:3626/foam/gapi/1 of-i2=,https://foam.net.internet2.edu:3626/foam/gapi/1 of-rutgers=,https://nox.orbit-lab.org:3626/foam/gapi/1 pg2=,https://www.emulab.net:12369/protogeni/xmlrpc/am/2.0 plc-bbn=,http://myplc.gpolab.bbn.com:12346/ plc-clemson=,http://myplc.clemson.edu:12346/
Interoperability Experiment
The following command were issued to create the slice used for this interoperability experiment:
$ omny.py createslice interop
ExoGENI Resources
The following sliver was created to reserve the 2 BBN ExoGENI VMs on shared VLAN 1750:
$ omny.py createsliver interop -a exobbn interop-2vm-shared-vlan-exo-bbn.rspec
The RSpec used: interop-2vm-shared-vlan-exo-bbn.rspec
The following sliver was created to reserved the 2 RENCI ExoGENI VMs on shared VLAN 1750:
$ omny.py createsliver interop -a exorci interop-2vm-shared-vlan-exo-rci.rspec
The RSpec used: interop-2vm-shared-vlan-exo-rci.rspec
The following sliver was created on the BBN ExoGENI FOAM aggregate to allow both the rack VMs and the local campus resources to connect to VLAN 1750:
$ omny.py createsliver interop -a of-exobbn interop-openflow-exobbn.rspec
The RSpec used: interop-openflow-exobbn.rspec
The following sliver was created on the RENCI ExoGENI FOAM aggregate to allow the rack VMs to connect to VLAN 1750:
$ omny.py createsliver interop -a of-exorci interop-openflow-exorci.rspec
The RSpec used: interop-openflow-exorci.rspec
InstaGENI Resources
The following sliver was created to reserve the 2 Utah InstaGENI VMs on shared VLAN 1750:
$ omny.py createsliver interop -a ig interop-2vm-shared-vlan-insta-utah.rspec
The RSpec used: interop-2vm-shared-vlan-insta-utah.rspec
The following sliver was created to reserve one Utah PG VM on shared VLAN 1750:
$ omny.py createsliver interop -a pg interop-1vm-shared-vlan-pg.rspec
The RSpec used: interop-1vm-shared-vlan-pg.rspec
The following sliver was created on the InstaGENI FOAM aggregate to allow both the rack VMs and the local campus resources to connect to VLAN 1750:
$ omny.py createsliver interop -a of-ig interop-openflow-insta-utah.rspec
The RSpec used: interop-openflow-insta-utah.rspec
Meso-scale Resources
The following slivers were created get compute resources at each of the Meso-scale aggregate mangers (Rutgers, BBN and Clemson):
$ omny.py createsliver interop -a pg2 --api-version 2 -T GENI 3 interop-wapg-rutgers.rspec $ omny.py createsliver interop -a plc-bbn interop-myplc-bbn.rspec $ omny.py createsliver interop -a plc-clemson interop-myplc-clemson.rspec
The RSpecs used: interop-wapg-rutgers.rspec interop-myplc-bbn.rspec interop-myplc-clemson.rspec
The following slivers were created at each of the Meso-scale FOAM aggregate (Rutgers, BBN and Clemson):
$ omny.py createsliver interop -a of-bbn interop-openflow-meso-bbn.rspec $ omny.py createsliver interop -a of-clemson interop-openflow-meso-clemson.rspec $ omny.py createsliver interop -a of-rutgers interop-openflow-meso-rutgers.rspec
The RSpecs used: interop-openflow-meso-bbn.rspec interop-openflow-meso-clemson.rspec interop-openflow-meso-rutgers.rspec
The following slivers were created at NRL and Internet2 FOAM Aggregate for VLAN 3716, which is used by all hosts in this experiment to exchange traffic:
$ omny.py createsliver interop -a of-i2 interop-openflow-meso-i2.rspec $ omny.py createsliver interop -a of-nlr interop-openflow-meso-nlr.rspec
The RSpecs used: interop-openflow-meso-i2.rspec interop-openflow-meso-nlr.rspec
Run Experiment
Determine which ExoGENI VMs have been assigned to you:
$ omny.py listrsources interop -a exobbn -o $ omny.py listresources interop -a exorci -o
Determine which InstaGENI Utah rack VMs have been assigned to you:
$ omny.py sliverstatus interop -a ig -o
Determine which PG Utah site VM was associated with VLAN 1750:
$ omny.py sliverstatus interop -a pg -o
Determine which hosts have been assigned to you for each of the meso-scale sites:
$ omny.py sliverstatus interop -a pg2 --api-version 2 -T GENI 3 -o $ omny.py sliverstatus interop -a plc-bbn -o $ omny.py sliverstatus interop -a plc-clemson -o
The output files created by the listresources and sliverstatus commands in this section include a "hostname" for each allocated compute resource. Additionally for the PG and InstaGENI VMs there is a port number required to connect.
Using information determined from the listresources and sliverstatus commands was able to login into each compute resource and exchange traffic between all Meso-scale, InstaGENI and ExoGENI compute resources, including the campus resources which were pass through the racks OF Switch.
Attachments (16)
- interop-1vm-shared-vlan-pg.rspec (1.0 KB) - added by 12 years ago.
- interop-2vm-shared-vlan-exo-bbn.rspec (1.6 KB) - added by 12 years ago.
- interop-2vm-shared-vlan-exo-rci.rspec (1.6 KB) - added by 12 years ago.
- interop-2vm-shared-vlan-insta-utah.rspec (1.4 KB) - added by 12 years ago.
- interop-myplc-bbn.rspec (831 bytes) - added by 12 years ago.
- interop-myplc-clemson.rspec (824 bytes) - added by 12 years ago.
- interop-openflow-insta-utah.rspec (1.4 KB) - added by 12 years ago.
- interop-openflow-meso-clemson.rspec (1.4 KB) - added by 12 years ago.
- interop-openflow-meso-i2.rspec (3.8 KB) - added by 12 years ago.
- interop-openflow-meso-nlr.rspec (2.8 KB) - added by 12 years ago.
- interop-wapg-rutgers.rspec (893 bytes) - added by 12 years ago.
- Mesoscale-GENIRacks-interop.jpg (278.6 KB) - added by 12 years ago.
- interop-openflow-meso-bbn.rspec (2.0 KB) - added by 12 years ago.
- interop-openflow-exobbn.rspec (1.5 KB) - added by 12 years ago.
- interop-openflow-exorci.rspec (1.5 KB) - added by 12 years ago.
- interop-openflow-meso-rutgers.rspec (1.6 KB) - added by 12 years ago.
Download all attachments as: .zip