| Version 26 (modified by , 12 years ago) (diff) |
|---|
IG-EXP-5: InstaGENI Network Resources Acceptance Test
This page captures status for the test case IG-EXP-5, which verifies the ability to support OpenFlow operations and integration with meso-scale compute resources and other compute resources external to the InstaGENI rack. For overall status see the InstaGENI Acceptance Test Status page.
Last Update: 08/24/12
Test Status
This section captures the status for each step in the acceptance test plan.
| Step | State | Ticket | Comments |
| Step 1 | Color(yellow,Complete)? | Experiment modified to use Utah rack | |
| Step 2 | Color(yellow,Complete)? | ||
| Step 3 | Color(yellow,Complete)? | ||
| Step 4 | Color(yellow,Complete)? | ||
| Step 5 | Color(yellow,Complete)? | ||
| Step 6 | Color(yellow,Complete)? | ||
| Step 7 | Color(yellow,Complete)? | ||
| Step 8 | Color(yellow,Complete)? | ||
| Step 9 | Color(yellow,Complete)? | ||
| Step 10 | Color(yellow,Complete)? | ||
| Step 11 | Color(yellow,Complete)? | ||
| Step 12 | Color(yellow,Complete)? | ||
| Step 13 | Color(#63B8FF,In Progress)? | ||
| Step 14 | Color(yellow,Complete)? | ||
| Step 15 | Color(yellow,Complete)? | ||
| Step 16 | Color(yellow,Complete)? | ||
| Step 17 | Color(yellow,Complete)? | ||
| Step 18 | Color(yellow,Complete)? | ||
| Step 19 | Color(yellow,Complete)? | ||
| Step 20 | Color(yellow,Complete)? | ||
| Step 21 | Color(yellow,Complete)? | ||
| Step 22 | Color(yellow,Complete)? | ||
| Step 23 | Color(yellow,Complete)? | ||
| Step 24 | Color(yellow,Complete)? | ||
| Step 25 | Color(yellow,Complete)? | ||
| Step 26 | Color(#63B8FF,In Progress)? | ||
| Step 27 | Color(#63B8FF,In Progress)? | ||
| Step 28 | Color(yellow,Complete)? | ||
| Step 29 | Color(yellow,Complete)? |
| State Legend | Description |
| Color(green,Pass)? | Test completed and met all criteria |
| Color(#98FB98,Pass: most criteria)? | Test completed and met most criteria. Exceptions documented |
| Color(red,Fail)? | Test completed and failed to meet criteria. |
| Color(yellow,Complete)? | Test completed but will require re-execution due to expected changes |
| Color(orange,Blocked)? | Blocked by ticketed issue(s). |
| Color(#63B8FF,In Progress)? | Currently under test. |
Test Plan Steps
This procedure are executed at Utah InstaGENI, not as originally planned at BBN additionally, due to rack delivery delays. Additionally, the initial run through for this procedure is also modified to use one set of user credentials, running two slices for one experiment.
The following aggregate managers nick_names are define in the omni_config used for this test:
ig-utah=,http://utah.geniracks.net/protogeni/xmlrpc/am pg=,http://www.emulab.net/protogeni/xmlrpc/am pg2=,https://www.emulab.net:12369/protogeni/xmlrpc/am/2.0 of-bbn=,https://foam.gpolab.bbn.com:3626/foam/gapi/1 of-clemson=,https://foam.clemson.edu:3626/foam/gapi/1 of-i2=,https://foam.net.internet2.edu:3626/foam/gapi/1 of-ig=,https://foam.utah.geniracks.net:3626/foam/gapi/1 of-rutgers=,https://nox.orbit-lab.org:3626/foam/gapi/1 plc-bbn=,http://myplc.gpolab.bbn.com:12346/ plc-clemson=,http://myplc.clemson.edu:12346/
1. As Experimenter1 (lnevers@bbn.com), determine PG site compute resources and define RSpec
Collect list resources from InstaGENI compute and network aggregate managers.
$ omni.py -a ig-utah listresources -o $ omni.py -a pg listresources -o $ omni.py -a of-ig listresources -o $ omni.py -a of-i2 listresources -o
Defined the IG-EXP-5-exp1-pg-site.rspec file to request the ProtoGENI site compute resources request.
2. Determine remote meso-scale compute resources and define RSpec
The RSpecs for experiment 1 are defined for the meso-scale sites at Rutgers, where one WAPG node (pg51) is used via Internet2
3. Define a request RSpec for OpenFlow network resources at the InstaGENI AM
Defined the RSpec IG-EXP-5-exp1-openflow-ig.rspec for the Utah InstaGENI FOAM Aggregate to allow the site PG site compute resources to access the core VLAN 3716.
4. Define a request RSpec for OpenFlow network resources at the remote I2 Meso-scale site
RSpecs were defined for Rutgers FOAM aggregate and WAPG compute resource. The following RSpecs were used for the meso-scale site:
- IG-EXP-5-exp1-rutgers-wapg.rspec - Rutgers WAPG nodes pg51 compute resource request RSpec.
- IG-EXP-5-exp1-openflow-rutgers.rspec - Rutgers FOAM Aggregate network resource request RSpec.
5. Define a request RSpec for the OpenFlow Core resources
The file IG-EXP-5-exp1-openflow-i2.rspec defines the Internet2 Core FOAM Aggregate network resources request RSpec.
6. Create the first slice
Created the first slice as with GPO ProtoGENI credentials:
$ omni.py createslice IG-EXP-5-exp1
7. Create a sliver for the BBN compute resources
Created slivers at the PG Utah compute resource aggregate
$ omni.py -a pg createsliver IG-EXP-5-exp1 IG-EXP-5-exp1-pg-site.rspec
8. Create a sliver at the I2 meso-scale site using FOAM at site
Created a sliver at the Rutgers FOAM for VLAN 3716:
$ omni.py -a of-rutgers createsliver IG-EXP-5-exp1 IG-EXP-5-exp1-openflow-rutgers.rspec
9. Create a sliver at of the Utah InstaGENI AM
Created slivers at the InstaGENI rack FOAM network resource aggregate
$ omni.py -a of-ig createsliver IG-EXP-5-exp1 IG-EXP-5-exp1-openflow-ig.rspec
10. Create a sliver for the OpenFlow resources in the core
Created slivers at Internet2 and NLR FOAM network resource aggregate:
$ omni.py -a of-i2 createsliver IG-EXP-5-exp1 G-EXP-5-exp1-openflow-i2.rspec
10a. Create a sliver for all remaining Meso-scale compute and network resources
WAPG nodes are part of the Utah PG aggregate, therefore a second request to add a resource does not work. Also modify sliver is not an available feature at this time, so this step is modified to create a second slice for experiment1. In order to create a second request at the PG site the following slice and sliver were created for the Rutgers WAPG node:
$ omni.py createslice IG-EXP-5-exp1a $ omni.py -a pg2 createsliver IG-EXP-5-exp1a --api-version 2 -t GENI 3 IG-EXP-5-exp1-rutgers-wapg.rspec
11. Log in to each of the compute resources and send traffic to the other end-point
The nodes assigned for compute resources were determined as follows:
$ omni.py -a pg sliverstatus IG-EXP-5-exp1 --api-version 2 -t GENI 3 -o $ omni.py -a pg2 sliverstatus IG-EXP-5-exp1a -o
The above created output files from which the assigned hosts can be determined:
$ egrep "hostname" IG-EXP-5-sliverstatus-SITENAME.json
Or you me also use the gcf/examples/readyToLogin.py script:
$ ./examples/readyToLogin.py -a pg IG-EXP-5-exp1 -o ... ================================================================================ Aggregate [http://www.emulab.net/protogeni/xmlrpc/am] has a ProtoGENI sliver. pc535.emulab.net's geni_status is: ready Login using: xterm -e ssh -i ~/.ssh/id_rsa lnevers@pc535.emulab.net -p 35642 & ================================================================================
12. Verify that traffic is delivered to target
Logged in with the following:
$ xterm -e ssh -i ~/.ssh/id_rsa lnevers@pc535.emulab.net -p 35642 &
Exchanged traffic with the remote meso-scale resource at Rutgers:
13. Review baseline, GMOC, and meso-scale monitoring statistics
Checking of monitoring status is replaced by iperf tests. Add capture here.
14. As Experimenter2, determine Utah site compute resources and define RSpec
As Experimenter2 (lnevers2@bbn.com) defined XXXX
15. Determine remote meso-scale NLR compute resources and define RSpec
The Clemson site was used as a remote meso-scale site and the file IG-EXP-5-myplc-clemson.rspec captures the Clemson MyPLC node planetlab4 compute resource request.
16. Define a request RSpec for OpenFlow network resources at the Utah InstaGENI AM
Defined the IG-EXP-5-exp2-pg-site.rspec file to request the ProtoGENI site compute resources request for experiment2.
17. Define a request RSpec for OpenFlow network resources at the remote NLR Meso-scale site
Defined the file IG-EXP-5-openflow-clemson.rspec to capture the Clemson FOAM Aggregate network resource request needed to allow the MyPLC node (planetlab4.clemson.edu) access to the OpenFlow network core.
18. Define a request RSpec for the OpenFlow Core resources
Defined IG-EXP-5-openflow-nlr.rspec to capture the NLR Core FOAM Aggregate network resources request.
19. Create the second slice
As experimenter2 (lnevers2@bbn.com) created a slice:
$ omni.py createslice IG-EXP-5-exp2
20. Create a sliver for the Utah compute resources
Created a sliver for the site resources at the Utah PG as follows:
$ omni.py -a pg createsliver IG-EXP-5-exp2 IG-EXP-5-exp2-pg-site.rspec
21. Create a sliver at the meso-scale site using FOAM at site
Created a sliver at the Clemson FOAM aggregate requesting the network resources required to allow the MyPLC host to access the OpenFlow Backbone VLAN 3716:
$ omni.py -a of-clemson createsliver IG-EXP-5-exp2 IG-EXP-5-exp2-openflow-clemson.rspec
22. Create a sliver at of the Utah InstaGENI AM
Created slivers at the InstaGENI rack FOAM network resource aggregate to allow the PG site resources to access the OpenFlow backbone:
$ omni.py -a of-ig createsliver IG-EXP-5-exp2 IG-EXP-5-exp2-openflow-ig.rspec
23. Create a sliver for the OpenFlow resources in the core
Created sliver at NLR FOAM network resource aggregates:
$ omni.py -a of-nlr createsliver IG-EXP-5-exp2 IG-EXP-5-exp2-openflow-nlr.rspec
24. Create a sliver for the meso-scale compute resources
Requested the MyPLC node at the Clemson resource aggregate:
$ omni.py -a plc-clemson createsliver IG-EXP-5-exp2 IG-EXP-5-exp2-myplc-clemson.rspec
25. Log in to each of the compute resources and send traffic to the other endpoint
Verify the status for each compute resource sliver, and use login information when ready:
$ omni.py -a plc-clemson sliverstatus IG-EXP-5-exp1 -o $ omni.py -a pg sliverstatus IG-EXP-5-exp1 -o
Used the readToLogin.py script to determine state and login:
$ ./examples/readyToLogin.py -a pg IG-EXP-5-exp2 .... ================================================================================ Aggregate [http://www.emulab.net/protogeni/xmlrpc/am] has a ProtoGENI sliver. pc496.emulab.net's geni_status is: notready Login using: xterm -e ssh -i /home/lnevers2/.ssh/geni_key lnevers2@pc496.emulab.net -p 35898 & ================================================================================ $ ./examples/readyToLogin.py -a plc-clemson IG-EXP-5-exp2 ... ================================================================================ Aggregate [http://myplc.clemson.edu:12346/] has a PlanetLab sliver. planetlab4.clemson.edu's geni_status is: ready (pl_boot_state:boot) Login using: xterm -e ssh -i /home/lnevers2/.ssh/geni_key pgenigpolabbbncom_IGEXP5exp2@planetlab4.clemson.edu & ================================================================================
Was able to exchange ping traffic between each of the compute resources.
At PG site resource:
[lnevers2@utah-pg ~]$ ping 10.42.11.104
At Clemson MyPLC site:
[lnevers2@utah-pg ~]$ ping 10.42.11.104
26. As Experimenter2, insert flowmods and send packet-outs only for traffic assigned to the slivers
Need to get controller that cand do flowmods.
27. Verify that traffic is delivered to target according to the flowmods settings
See step 26.
28. Review baseline, GMOC, and monitoring statistics
Insert Iperf results here:
Reviewed the monitoring information available about this experiment, first found the sliver in the list of slices:
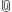 The selected the detail panel:
The selected the detail panel:
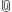
Then selected the sliver resources panel:
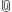
Also checked the sliver measurements:
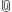
29. Stop traffic and delete slivers
Stopped traffic, and deleted slivers.
As Expeirenter1:
$ omni.py -a pg deletesliver IG-EXP-5-exp1 $ omni.py -a of-rutgers deletesliver IG-EXP-5-exp1 $ omni.py -a of-ig deletesliver IG-EXP-5-exp1 $ omni.py -a of-i2 deletesliver IG-EXP-5-exp1 $ omni.py -a pg2 deletesliver IG-EXP-5-exp1a
As Expeirenter2:
$ omni.py -a pg deletesliver IG-EXP-5-exp2 $ omni.py -a of-clemson deletesliver IG-EXP-5-exp2 $ omni.py -a of-ig deletesliver IG-EXP-5-exp2 $ omni.py -a of-nlr deletesliver IG-EXP-5-exp2 $ omni.py -a plc-clemson deletesliver IG-EXP-5-exp2
Attachments (7)
- IG-EXP-5-exp1-sliver-detail-atGPO.jpg (266.8 KB) - added by 11 years ago.
- IG-EXP-5-exp1-slivers.jpg (745.8 KB) - added by 11 years ago.
- IG-EXP-5-exp1-stats.jpg (317.9 KB) - added by 11 years ago.
- IG-EXP-5-exp1.jpg (126.8 KB) - added by 11 years ago.
- IG-EXP-5-exp2-slivers.jpg (475.0 KB) - added by 11 years ago.
- IG-EXP-5-exp2.jpg (128.8 KB) - added by 11 years ago.
- IG-EXP-5-slices.jpg (156.4 KB) - added by 11 years ago.
