Understanding the AM API using Content Centric Networking
1. Design the Experiment
|
2. Establish the Environment
2.1 Set up ssh keys and configure Omni
If your account is from the GENI Portal follow the instructions here.
If your account is from the iMinds member authority (https://www.wall2.ilabt.iminds.be) follow the instructions here.
3. Obtain Resources
3.1 Create a slice
Create a slice using omni and the slice name of your choice. From now on that slice name will be referred to as SLICENAME.
$ omni createslice SLICENAME
3.2. Load a simple topology in jFed
For this exercise, we will edit an existing RSpec file. Start by loading this predefined topology into jFed.
|
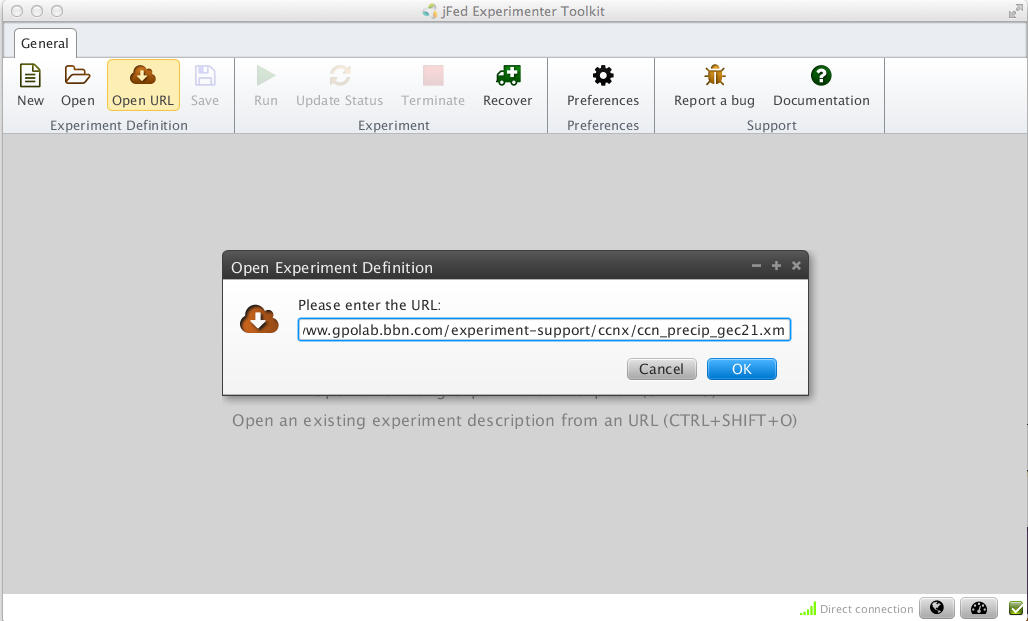
Figure 3-1 Import an RSpec into jFed. |
|
3.3. Modify the RSpec to automatically install and execute CCNX software
For this experiment, we need to install the following software on the nodes:
- The CCNX software (ccnx-0.6.2.tar.gz)
- Scripts that set up the CCNX software (ccnx-setup.tar.gz)
- Scripts used to pull atmospheric precipitation data using the CCNX protocol (ccnx-atmos.tar.gz)
When the nodes start up, we need the following scripts to be executed:
- Script that sets up the node (node-setup)
- Script that sets up the ccnx protocol (ccnx-setup)
- Script that setup up ccnx protocol routes (add-precip-routes)
We automate the installation and running of the proper software using install and execute scripts in the RSpec. These can be added by double clicking on a node, and then clicking a "+" under the "Boot scripts" tab.
|
You DO NOT have to specify install and execute scripts for the nodes as they have already been done for you. You can check this by clicking on the icons for these nodes.
In general, be very careful when entering this information. These commands will not be executed until the nodes boot up and may fail silently. |
|

Figure 3-2 Edit all three nodes |
|

Figure 3-3 Copy the router and user nodes. |
|

Figure 3-4 Draw links to connect the nodes. |
|

Figure 3-5 Edit the next hop on router1 and user0. |
3.4. Export the modified request RSpec
Now we will pull back some of the covers and inspect exactly what jFed has been doing for us when preparing the RSpecs for the experiments we design. Each node and link has a corresponding element in the RSpec, and the details of the component configuration (such as the install and execute services we requested above) are specified with attributes, or sometimes child elements, within those portions of the document.
|
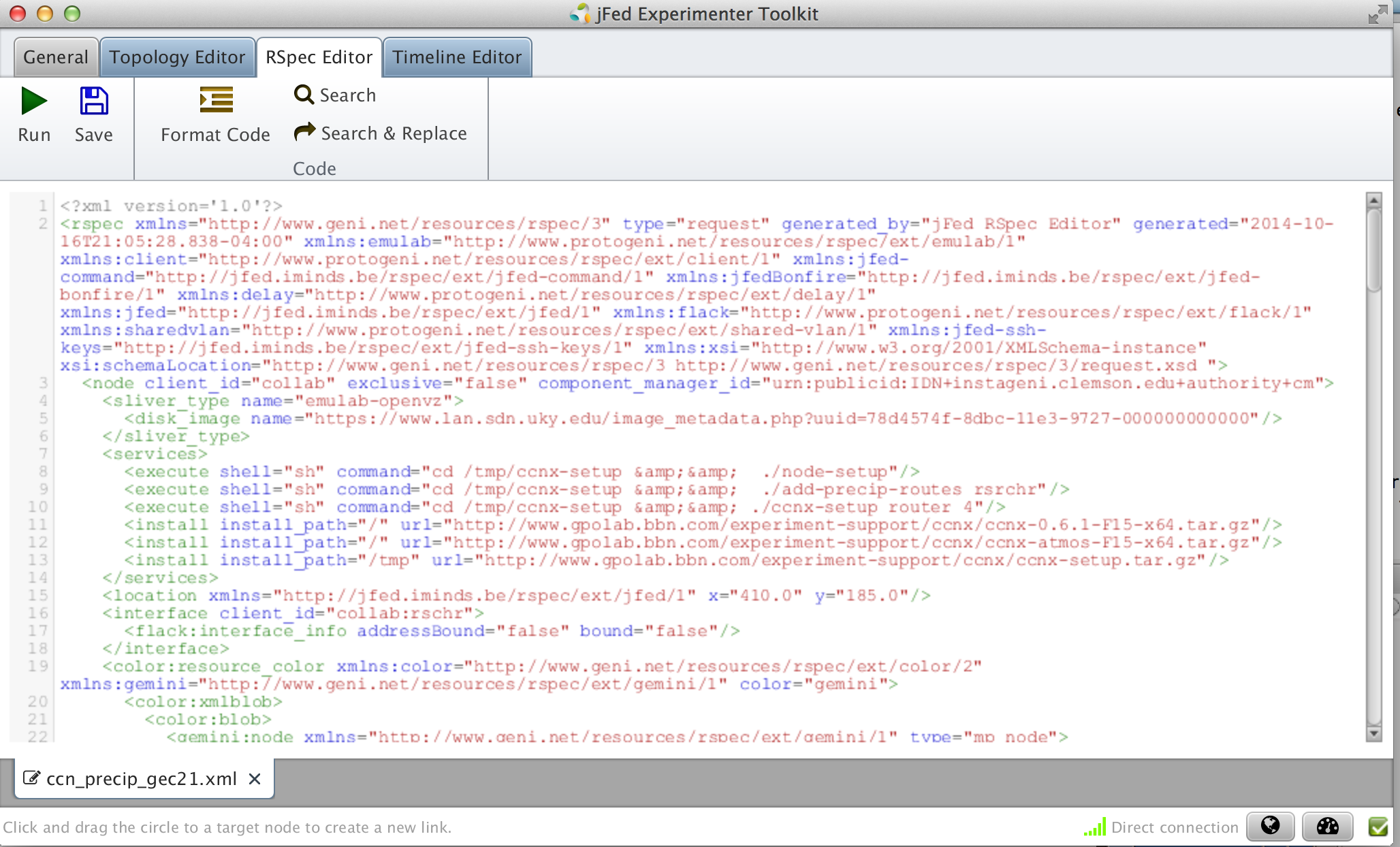
Figure 3-5 View the final request RSpec |
|
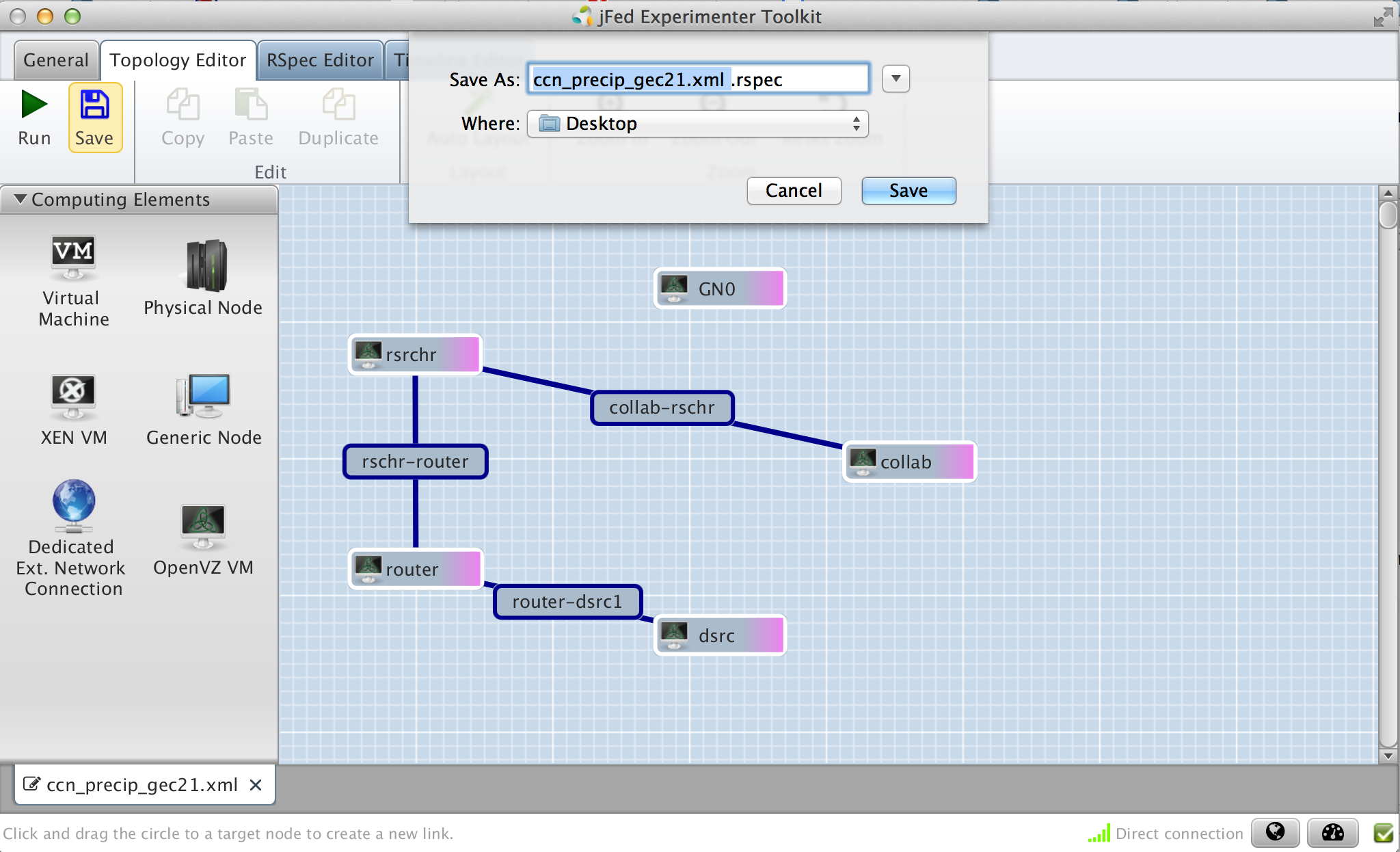
Figure 3-6 Save the final request RSpec |
3.6. Instantiate the new experiment using Omni
For this step, we'll change the approach a bit and switch to a new client tool, the command line Omni client.
From a terminal, please enter the command:
$ omni -a AM_NICKNAME createsliver SLICENAME RSPEC_FILE
where AM_NICKNAME is the nickname for your assigned aggregate manager and SLICENAME is the name of the slice you created earlier (both of these are given on your worksheet). RSPEC_FILE should be replaced with the filename of the RSpec you saved in step 4.
If all is well, Omni should give you a number of informational messages, such as:
INFO:omni:Loading config file /home/geni/.gcf/omni_config
It should quickly proceed to the point where it makes the request to the remote manager:
INFO:omni:Creating sliver(s) from rspec file /home/geni/Downloads/experiments.rspec for slice ...
This step can sometimes be time-consuming, so please be patient. If it succeeds, within a couple of minutes Omni should report:
INFO:omni: Completed createsliver:
and your resource reservation is complete!