1. Design the Experiment
- In today's experiment you will use resources at the aggregate assigned to you by the tutorial leader
|
2. Establish the Environment
If your account is from the GENI Portal follow the instructions in the Section 2 of this exercise.
If your account is from the iMinds member authority, follow the instructions at ...
3. Obtain Resources
3.2. Load the experiment topology
For this exercise, we will edit an existing RSpec file. Start by loading this predefined topology into jFed.
- Launch jFed.
- Click on the Open URL menu (see figure)
- Enter the url http://www.gpolab.bbn.com/experiment-support/iMinds/rspec-ccn-fedora-eu.rspec .
|
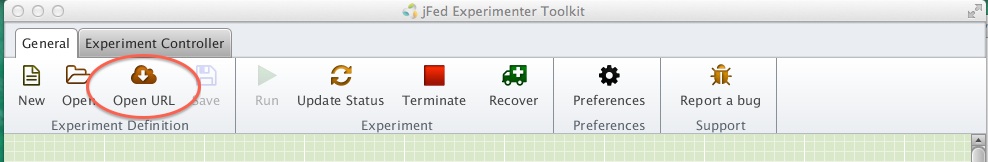
Figure 3-1 Import an RSpec into jFed.
|
- You should see a 4-node linear topology. Double-click on each node to make sure you are using the aggregate assigned to you ( iMinds Virtual Wall 1 or iMinds Virtual Wall 2 . If needed, change the aggregate from the drop-down list and click Save. Do this for all nodes
|

Figure 3-1 Make sure resource is being requested from the correct aggregate.
|
=== 3.3. Modify the RSpec to automatically install and execute CCNX software ===
For this experiment, we need to install the following software on the nodes:
* The CCNX software (ccnx-0.6.1-F15-x64.tar.gz)
* Scripts that set up the CCNX software (ccnx-setup.tar.gz)
* Scripts used to pull atmospheric precipitation data using the CCNX protocol (ccnx-atmos-F15-x64.tar.gz)
When the nodes start up, we need the following scripts to be executed:
* Script that sets up the node (node-setup)
* Script that sets up the ccnx protocol (ccnx-setup)
* Script that setup up ccnx protocol routes (add-precip-routes)
We automate the installation and running of the software using install and execute scripts in the RSpec.
{{{
#!html
- Click on the the i button on the collab node (see figure) to manipulate details of the node configuration.

|
When you open the node information, you should see options like those
in the figure.
|
|
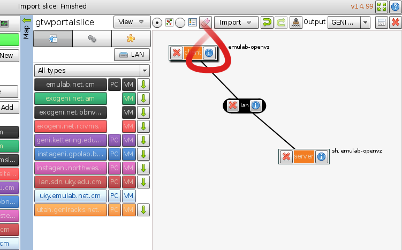
Figure 3-2 Edit the nodes
|
- The way we will request installation of the proper software and execution
of our experiment is to add install and execute services.
|
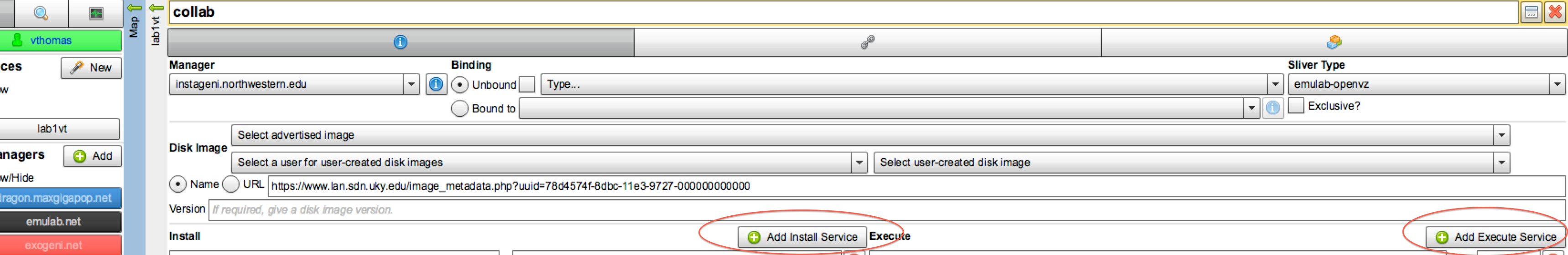
Figure 3-3 Specify the install and execute scripts
|
-
First, select "+ Add Install Service" as shown. In the box the Install box enter (cut-and-paste):
http://www.gpolab.bbn.com/experiment-support/ccnx/ccnx-0.6.1-F15-x64.tar.gz
and enter
/
in the box next to it (labeled 'in').
Now, select "+ Add Install Service" again. This time add:
http://www.gpolab.bbn.com/experiment-support/ccnx/ccnx-atmos-F15-x64.tar.gz
and enter
/
in the box next to it (labeled 'in').
Similarly, add a third install service:
http://www.gpolab.bbn.com/experiment-support/ccnx/ccnx-setup.tar.gz
and enter
/tmp
in the box next to it (labeled 'in'). Note the different install directory this time.
- In a similar manner, use the "+ Add Execute Service" button to add three execute services on this node:
cd /tmp/ccnx-setup && ./node-setup
cd /tmp/ccnx-setup && ./add-precip-routes rsrchr
cd /tmp/ccnx-setup && ./ccnx-setup router 4
 | Be very careful when entering this information -- these commands will not
be executed yet, so it will be some time before you will see any relevant
error messages if there is a mistake here. |
|
- Once both the install and execute services are specified, click Apply.
- You DO NOT have to specify install and execute scripts for the other nodes as they have already been done for you. You can check this by clicking on the i button next to the icons for these nodes.
|
}}}
=== 3.4. Add GEMINI extensions ===
This assignment has the option of instrumenting the experiment and viewing graphs of traffic on links using the GENI Desktop. To prepare for this option, you need to turn on GEMINI extensions. Do this by
{{{
#!html
- Select the tab in the middle panel of Flack with the 'puzzle piece' icon
- Check the box "Use GEMINI Extensions"
|
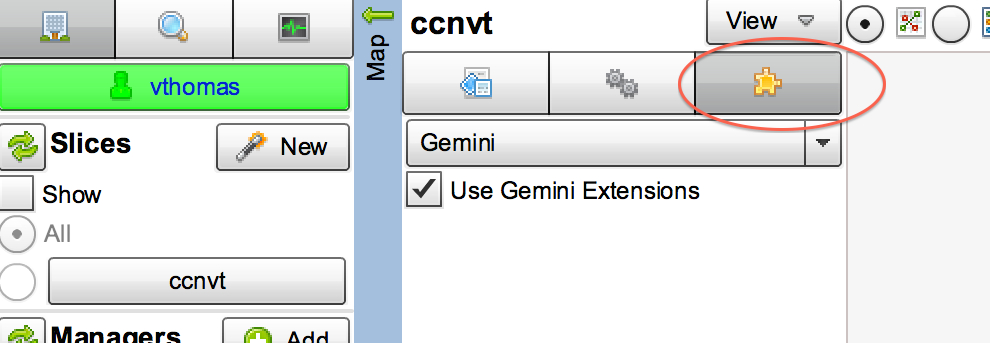
Figure 3-4 Add GEMINI Extensions
|
}}}
=== 3.5. Export the modified request RSpec ===
Now we will pull back some of the covers and inspect exactly what Flack has been doing for us when preparing the RSpecs for the experiments we design. Each node and link has a corresponding element in the RSpec, and the details of the component configuration (such as the install and execute services we requested above) are specified with attributes, or sometimes child elements, within those portions of the document.
{{{
#!html
- From the View menu (see figure), select the Preview
request document(s) item. This will bring up a window showing the
current RSpec -- please take a moment to inspect it. The XML elements
under the "flack" namespace were added for Flack's internal
use (containing information about the canvas layout, editing history, etc.),
and are ignored by aggregate managers. The `<node>` and `<link>` elements
contain the specification for the components we will request, and if you
look closely, you will be able to see the install and execute service
elements you added earlier.
|
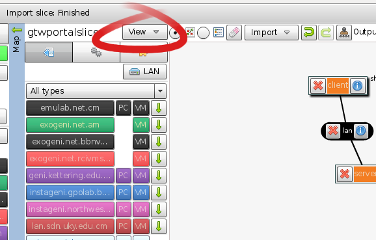
Figure 3-5 View and save the final request RSpec
|
- Use the Save to file button (in the upper left) to
a local copy of your RSpec with the name lab1_rspec.xml. We'll use this in the next step to
demonstrate how other client tools also use RSpec files to communicate
requests to aggregate managers.
|
}}}
=== 3.6. Instantiate the new experiment using Omni ===
For this step, we'll change the approach a bit and switch to a new client tool, the command line Omni client.
From a terminal, please enter the command:
{{{
$ omni -a AM_NICKNAME createsliver SLICENAME RSPEC_FILE
}}}
where {{{AM_NICKNAME}}} is the nickname for your assigned aggregate manager and {{{SLICENAME}}} is the name of the slice you created earlier (both of these are given on your worksheet). {{{RSPEC_FILE}}} should be replaced with the filename of the RSpec you saved in step 4.
If all is well, Omni should give you a number of informational messages, such as:
{{{
INFO:omni:Loading config file /home/geni/.gcf/omni_config
}}}
It should quickly proceed to the point where it makes the request to the remote manager:
{{{
INFO:omni:Creating sliver(s) from rspec file /home/geni/Downloads/experiments.rspec for slice ...
}}}
This step can sometimes be time-consuming, so please be patient. If it succeeds, within a couple of minutes Omni should report:
{{{
INFO:omni: Completed createsliver:
}}}
and your resource reservation is complete!
----
= [wiki:GENIExperimenter/Tutorials/GettingStarted_PartII_ccn/Procedure/Setup Introduction] =
= [http://groups.geni.net/geni/wiki/GENIExperimenter/Tutorials/GettingStarted_PartII_ccn/Procedure/Execute Next: Execute] =