| Version 9 (modified by , 10 years ago) (diff) |
|---|
GENI Inter-Aggregate Stitching Tutorial
Step 1: Download and Import Initial Resource Request RSpec
This tutorial starts with an existing RSpec which includes 3 nodes in a linear topology. The resources are un-bound, which means the resources are not affiliated with any rack aggregate.
To get started download the initial RSpec file and import it into the flack to make modification as follows:
1. Download the Rspec named 3nodes-1rack.rspec to your local system
2. Login to the GENI Portal at https://portal.geni.net
3. Go to the Projects panel and click on the "Create Slice" button for the tutorial project. (If you are running this tutorial on your own, you may use any existing project.) On the resulting page named "Create New Slice" enter a slice name and click on "Create slice" button. For the purpose of this tutorial we will assume the slice is named MySlice, but you should use a unique name because each slice name must be unique within a project.
4. Once you are on the slice page (Home → Project MyProject → Slice MySlice) select the "Launch Flack" button. (You must wait for all aggregates to show up in the MySlice panel before you can proceed.
5. Select the pull-down named "Import" and choose "Import from file" option:
6. Navigate to the file location and choose select the 3nodes-1rack.rspec file.
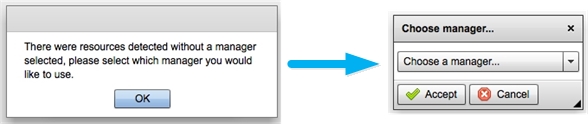
7. You will see a pop-up that states no manager is associated with the resource, click "OK" and another pop-up will have you select an aggregate manager, select one of the two aggregate manager assigned to you and click on "Accept" button. (If you are running this tutorial on your own, you may use any aggregate found on the GENI Network Stitching Sites page.
Step 2: Modify RSpec to include 2 Aggregates with stitched link
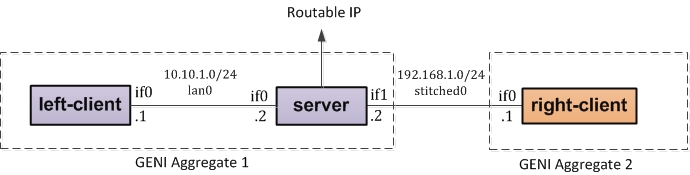
Now that you have imported the RSpec, we will modify it to include a stitched link. Modification in this section will result in the following topology:
To get to the above topology, we must make changes to the resources imported into flack. We will now modify "right-client" to become part of a different rack aggregate and then modify its link to a stitched link.
8. Click on the "right-client" VM information button .
.
9. In the "right-client" details panel look for the "Manager" pull-down. Modify the aggregate to the second manager manager that was assigned to you at the beginning of the tutorial, do not use the aggregate selected initially. Click on the "Apply" button. (If you are running this tutorial on your own, you may use another site aggregate found on the GENI Network Stitching Sites page.)
10. You will now see that the link from the "server" VM to "right-client" VM is now a gre-tunnel link. Select the gre-tunnel link information button to modify the link type to "stitched" and click "Apply" button.
to modify the link type to "stitched" and click "Apply" button.
11. You should now see that the topology has changed and that between the "server" VM and the "right-client" VM the link is now named "stitched0".
12. We will now export this topology to a file by selecting the "View" pull-down and choosing "Preview requested document(s)". You will now choose "Save to file" to export the RSpec we just modified. For the purpose of this experiment we will assume the file is saved to "MyTopology.rspec", but you can choose any name or location, just make a not of it.
We are now ready to use stitcher to create the stitched topology for our experiment!
Step 3: Run stitcher tool to get resources
The stitcher tools delivers the same features as the omni tool, but with one major difference, the stitcher will submit sliver request at all aggregate in your requests RSpec. Let's now set up slivers for the request RSpec generated in the previous section:
13. Use stitcher to create a sliver and using the slice that we created in flack. The request will use the RSpec that was saved to file in the setup steps:
$ stitcher createsliver MySlice MyTopology.rspec 11:48:18 INFO stitcher: Loading agg_nick_cache file '/Users/lnevers/.gcf/agg_nick_cache' 11:48:18 INFO stitcher: Loading config file /Users/lnevers/.gcf/omni_config 11:48:18 INFO stitcher: Setting option useSliceMembers based on omni_config setting 11:48:18 INFO stitcher: Using control framework portal 11:48:18 INFO stitcher: Member Authority is https://ch.geni.net/MA (from config) 11:48:18 INFO stitcher: Slice Authority is https://ch.geni.net/SA (from config) 11:48:19 INFO stitcher: Checking that slice MySlice is valid... 11:48:19 INFO stitcher: Slice urn:publicid:IDN+ch.geni.net:ln-prj+slice+MySlice expires on 2014-03-17 14:57:35 UTC 11:48:20 INFO stitcher: Stitched reservation will include resources from these aggregates: 11:48:20 INFO stitcher: <Aggregate wisconsin-ig> 11:48:20 INFO stitcher: <Aggregate stanford-ig> 11:48:20 INFO stitcher: <Aggregate ion> 11:48:20 INFO stitch.Aggregate: Stitcher doing createsliver at https://www.instageni.wisc.edu:12369/protogeni/xmlrpc/am 11:48:38 INFO stitch.Aggregate: Allocation at <Aggregate wisconsin-ig> complete. 11:48:38 INFO stitch.Aggregate: Stitcher doing createsliver at https://www.instageni.stanford.edu:12369/protogeni/xmlrpc/am 11:48:58 INFO stitch.Aggregate: Allocation at <Aggregate stanford-ig> complete. 11:48:58 INFO stitch.Aggregate: Stitcher doing createsliver at http://geni-am.net.internet2.edu:12346 11:49:43 INFO stitch.Aggregate: DCN AM <Aggregate ion>: must wait for status ready.... 11:49:43 INFO stitch.Aggregate: Pause 30 seconds to let circuit become ready... 11:50:20 INFO stitch.Aggregate: DCN circuit 74871 is ready 11:50:27 INFO stitch.Aggregate: Allocation at <Aggregate ion> complete. 11:50:27 INFO stitch.launcher: All aggregates are complete. 11:50:27 INFO stitcher: Saved combined reservation RSpec at 3 AMs to file MySlice-manifest-rspec-stitching-combined.xml Stitching success: Reserved resources in slice MySlice at 3 Aggregates (including 1 intermediate aggregate(s) not in the original request), creating 1 link(s).
We are now ready to run the experiment!