| Version 13 (modified by , 11 years ago) (diff) |
|---|
Obtain Resources: create a slice and reserve resources
Introduction: Getting Started with GENI and the GENI Portal
|
|

|
|

|
|
Instructions
Now that you are a member of a project, you can create a slice and reserve resources.
4. Create a slice
|
|
If you are following on from the Flack tutorial, and the slice you created then has not yet expired, you can reuse the same slice -- skip ahead to step 2 (Import a simple experiment). |
Otherwise, you will need to create a new slice, which can be done through the GENI Portal, as follows:
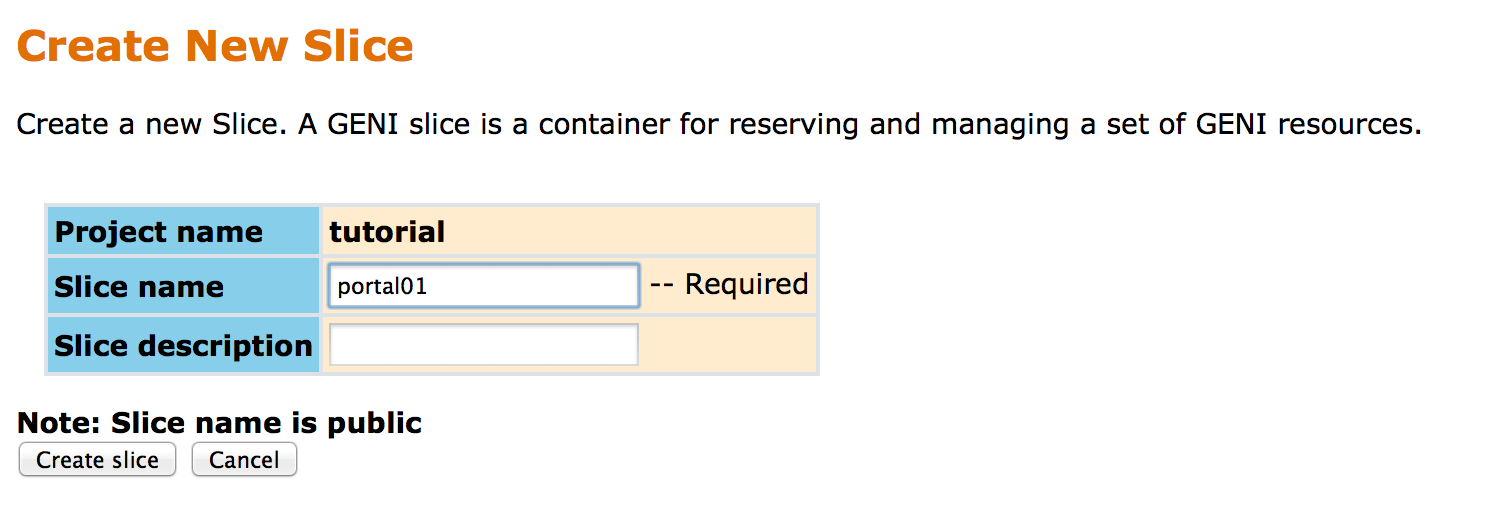
|
|
4.1. Import a simple experiment
Next, we will learn about the mechanism Flack had been using when bringing up slivers for us. Behind the scenes, slivers are described in GENI rspec files, which are relatively simple XML documents precisely describing the components, configurations, and topologies.
For this exercise, we will start with a prepared rspec file.
|
You should load this topology into Flack:
|
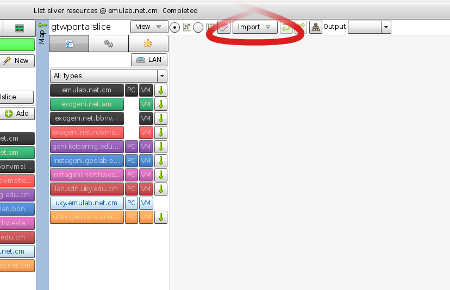 |
4.2. Modify the experiment to automatically install and execute network test software
1. For this sample experiment, we'd like some additional software (Apache, iperf, and a couple of custom scripts) loaded onto the VMs to perform traffic measurements. In Flack, you can manipulate the details of the request for a node or link with the "i" button on the relevant component on the canvas (see figure). Do this now on the client node. |
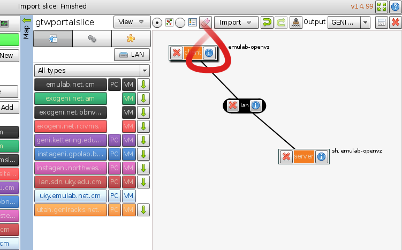 |
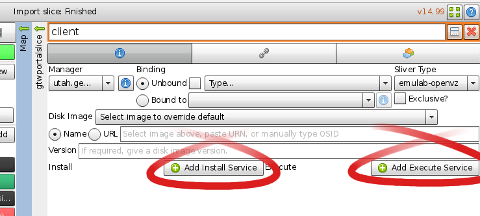
2. When you open the node information, you should see options like those in the figure. The controls available to you will vary depending on the types of the aggregate manager and the component, but many features are widely available throughout GENI. 3. The way we will request installation of the proper software and execution of our experiment is to add install and execute services. First, select "+ Add Install Service" as shown. Flack will now prompt you for an "Archive URL" and a location for installation. For this experiment, please enter the URL: http://protogeni.net/gec17tut2.tar.gzand ask for this to be installed in: /local 4. Similarly, please use the "+ Add Execute Service" button to request that the command: sudo /local/install-script.shbe executed using sh
5. Once both the "install" and "execute" services are specified, please choose "Apply". Then repeat the procedure to add the same "install" and "execute" services on the server node. If you have time, it would be instructive to download the "hellogeni-install" file yourself, and inspect the contents. You will see the install-script.sh file referred to in the execute service, and this is a good example of how you can ask for GENI components to perform tasks for you without any manual intervention. This is a very useful facility for large experiments! |
 |
4.3. Export the modified experiment
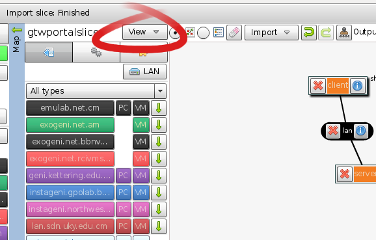
Now we will pull back some of the covers and inspect exactly what Flack has been doing for us when preparing the rspecs for the experiments we design. Each node and link has a corresponding element in the rspec, and the details of the component configuration (such as the install and execute services we requested above) are specified with attributes, or sometimes child elements, within those portions of the document.
|
 |
5. Instantiate the new experiment using Omni
For this step, we'll change the approach a bit and switch to a new client tool: the command line Omni client.
5.1. Configure Omni
- Go to your browser tab with the GENI Portal page.
- On that page, select the tab labeled "Profile". This is on the upper right of the page.
- Scroll down to the section labeled "Option 1: Automatic omni configuration".
- Click on "customized configuration data" in Step 2 of this section.
- Make sure your default project is "ICDCS13". If not, make that your default.
- Click "Download omni bundle" and "save" the file. The file is saved in your Downloads folder.
- Open a terminal window and type {{{omni-configure.py -f portal. The cert and key files you need will be installed in the appropriate folders.
5.2. Send the CreateSliver request to the AM
Once Omni is installed and configured, bringing up the sliver you designed earlier is a one-step procedure. Just enter the command:
$ omni.py -a AM_NICKNAME createsliver SLICENAME RSPEC_FILE
where AM_NICKNAME is the nickname for your assigned aggregate manager and SLICENAME is the name of the slice you created at the portal (both of these are given on your worksheet). RSPEC_FILE should be replaced with the filename of the rspec you saved in step 4.
If all is well, Omni should give you a number of informational messages, such as:
INFO:omni:Loading config file /home/geni/.gcf/omni_config
It should quickly proceed to the point where it makes the request to the remote manager:
INFO:omni:Creating sliver(s) from rspec file /home/geni/Downloads/experiments.rspec for slice ...
This step can sometimes be time-consuming, so please be patient. If it succeeds, within a few minutes Omni should report:
INFO:omni: Completed createsliver:
and your sliver is complete!
Next: Inspect the experiment execution
Attachments (5)
-
example.rspec (664 bytes) - added by 11 years ago.
A simple rspec with two nodes and a link.
- flack-import-small.png (40.1 KB) - added by 11 years ago.
- flack-node-info-small.png (42.0 KB) - added by 11 years ago.
- flack-services-small.png (43.3 KB) - added by 11 years ago.
- flack-view-small.png (42.8 KB) - added by 11 years ago.
Download all attachments as: .zip
