|
Step 1: Get Ready:
The first thing we need to do is login to the portal.
- Go to the GENI Experimenter Portal and click the Use GENI button.
- From the Drop Down menu select your institution. If you got an account through the GENI Identity Provider, please select GENI Project Office.
Tip: Start typing the name of your institution and see the list become smaller.
- You will be transferred to the Login Page of your institution. Fill in your username and password.
|
Step 2: Launch your experiment:
- At the portal home page click the +New slice button.
Tip: If you are not a member of any project and you don't know how to procede, email us
- Name your slice xxxfw (where xxx are your initials), since slice names within a project must be unique. If you like, enter a description of the slice (optional).
- Click Create slice .
- Once the slice page loads, click the Add Resources button.
NOTE: If you get a warning about not having uploaded ssh keys just follow the instructions on providing an ssh key before you proceed.
- Scroll down to the Choose RSpec section, and from the drop down list select the existing RSpec labeled OpenFlow Firewall.
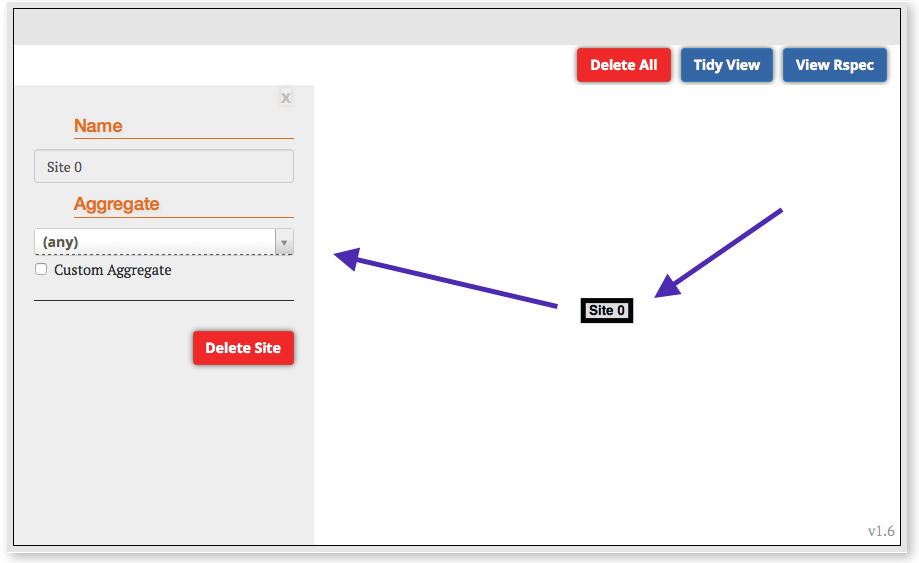
- Notice that a graphical representation of the proposed topology appears in the canvas above the Choose RSpec section. Click the Site 1 node, and then select any InstaGENI aggregate (any with InstaGENI in it's name) from the Site drop down list which appears on the left. You will need to choose an aggregate where you want this topology to be instantiated. Click on the Site 1 box and a panel on the left side of the canvas will appear. Choose any aggregate with InstaGENI in it's name.
- Click on the Reserve Resources button on them bottom left part of the screen.
- Repeat the above steps to create a second slice called xxxnat (where xxx are your initials) using the OpenFlow NAT RSpec
- Wait while your resources are being reserved. This will take several minutes so be patient. The nodes will turn green to signify that your resources are ready.
|
|
|




