| Version 51 (modified by , 11 years ago) (diff) |
|---|
This page will guide you through running a simple non-IP experiment in GENI, using the Omni command line tool. We are going to take advantage of the Layer 2 links between nodes and run a non-IP experiment. The only thing you will need is a GENI account. If you don't already have one, sign up!
1. Configure Omni with your GENI account
Omni is a command-line tool that will help you reserve resources in GENI. Follow these steps to download and configure Omni.

|
|
- Verify that you have the necessary credential and key files . Run:
ls ~/.ssh ~/.ssl
The output looks like :geni@geni-vm:~$ ls ~/.ssh ~/.ssl /home/geni/.ssh: config geni_key geni_key.pub /home/geni/.ssl: geni_cert_enc.pem geni_cert.pem
geni_cert.pem Cleartext certificate, i.e. does not require any passphrase geni_cert_enc.pem Encrypted certificate geni_key The private key that you will use to login to the nodes geni_key.pub The corresponding public key that will be uploaded to the nodes Note: Depending on the setup of your host you might see more files than the ones listed above.
- Optional - Look around the omni_config file
Open the file
~/.gcf/omni_configusing either vim or emacs. Close to the top of the file you will see two parameters calleddefault_cfandusers. Your username should be at least listed in the user section. Look for the sections in the file that are named[pg]and[<username>].
In the
[<username>]section, the information need for logging-in to reserved compute resources are provided. It includes your unique user URN and a public key that would be uploaded to the hosts that you reserve.
In the
[pg]section you configure Omni to use your personal information. The cert and the key attribute point to files that we have manually downloaded from pgeni.gpolab.bbn.com. This is equivalent to the Download action of Flack.
Another interesting section to look at is the
[aggregate-nicknames]sections. Flack already knows the URL for all the AMs and present you a list of AMs to choose from using a short, descriptive name. In Omni a user is required to pass the URL for each call to the GENI AM API. In this section the user gets a chance to provide short descriptive names to the URLs that are easier to memorize and use.
2. Launch your experiment
Now that you have configured Omni we are ready to design our experiment, which going over most of the Omni commands.
- Create a slice. The first thing to do when preparing to run a GENI experiment is to create a slice. Name your slice something like xxxomni (where xxx are your initials). Type
omni.py createslice <slicename>
- Verify that your slice was created. Use the
listmyslicescommand, of omni:omni.py listmyslices <username>
- Renew your slice. To extend the lifetime of your slice type:
omni.py renewslice <slicename> <YYYYMMDD>
- See available resources. For this experiment we are going to use the Aggregate manager of ProtoGENI in Utah. In order to see what each AM offers you can use the
listresourcescommand. Type:omni.py listresources -a pg-utah -o
The-ooption will save the output to a file. The filename is chosen by Omni and printed as part of the output. The output will look like :geni@geni-VirtualBox:~$ omni.py listresources -a pg-utah -o INFO:omni:Loading config file /home/geni/.gcf/omni_config INFO:omni:Using control framework pg INFO:omni:Saving output to a file. INFO:omni:Substituting AM nickname pg-utah with URL https://www.emulab.net/protogeni/xmlrpc/am/2.0, URN unspecified_AM_URN INFO:omni:Listed resources on 1 out of 1 possible aggregates. INFO:omni:Writing to 'rspec-www-emulab-net-protogeniv2.xml' INFO:omni: ------------------------------------------------------------ INFO:omni: Completed listresources: Options as run: aggregate: ['pg-utah'] framework: pg output: True Args: listresources Result Summary: Queried resources from 1 of 1 aggregate(s). Wrote rspecs from 1 aggregate(s) to 1 file(s) Saved listresources RSpec at 'unspecified_AM_URN' (url 'https://www.emulab.net/protogeni/xmlrpc/am/2.0') to file rspec-www-emulab-net-protogeniv2.xml; INFO:omni: ============================================================
In the last line of the output Omni will tell you the name of the file that output is saved at. In the example above this would berspec-www-emulab-net-protogeniv2.xml. Open the file that Omni saved and just take a look to see how an advertisement RSpec looks like. In order to see only available resources type:omni.py listresources -a pg-utah --available -o
- Reserve resources. To be able to reserve resources you will need to craft a request rspec. For this example we have created the rspec and made it available for you to use. Type:
omni.py createsliver -a pg-utah <slicename> http://www.gpolab.bbn.com/experiment-support/HelloGENI/hellogeni.rspec
- See the reserved resources. You can use the
listresourcescommand, to see what resources are reserved at an Aggregate.omni.py listresources -a pg-utah <slicename>
- Optional- Extend the lifetime of your reservation. The lifetime of your reservation can never exceed the lifetime of your slice and is usually set to a default value. For the purpose of this exercise you don't need to renew your reservation. But if you wanted to do this this is what the command would look like:
omni.py renewsliver -a pg-utah <slicename> <YYYYMMDD>
- Check the status of your resources. Type:
omni.py sliverstatus -a pg-utah <slicename>
Thesliverstatuscommand reports the status of your overall GENI slice. When the status is ready we are ready to continue to the next step.
3. View your results
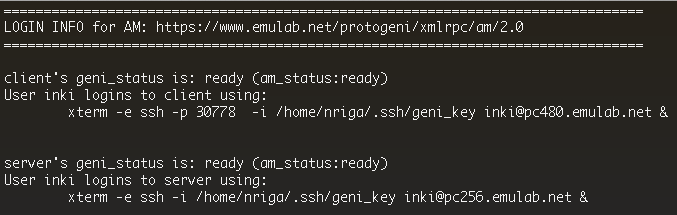
For this example experiment we used the install script facility to automatically install the necessary software and kick-off the experiment. In this very simple setup, we have installed and launched a web server as well as an iperf server, on the server host. On the client, we have started some processes to test both of these services. To view the results of this experiment:
|
|
|
The script will return the actual command that you would need to use for logging in. However we don't need this information right now. At the end of the script find the information that corresponds to the server host, and see what host was reserved (this will be the hostname after your username in the ssh command line, it should look like pcxxx.emulab.net ).
|
||

|
|
|
- Optional: Manually generate traffic While conducting experiments in GENI, you will often want to run commands directly on the nodes. In this optional step, you will log in to a node and issue commands directly to it.
- Follow these instructions and log in to the client node
- When you have successfully logged in, run this command
iperf -c server -P 2
This task shouldn't take more than 30 seconds. Change the number after the-Pargument and watch how the performance is affected while you change the number of parallel TCP connections. - Scroll all the way down the server iperf log, and look at the logs for your transfers
4. Cleanup
After you are done with your experiment, you should always release your resources so that other experimenters can use the resources.
- In the terminal, where you have been running your omni commands do:
omni.py deletesliver -a pg-utah <slicename>
