| Version 1 (modified by , 11 years ago) (diff) |
|---|
CCN ASSIGNMENT
In this tutorial, we will show how named data networking on GENI.
Overview
This Project leverages resources on the GENI aggregate in order to experiment with Content Centric Network.
Please remember to release the resources you when you are done with them.
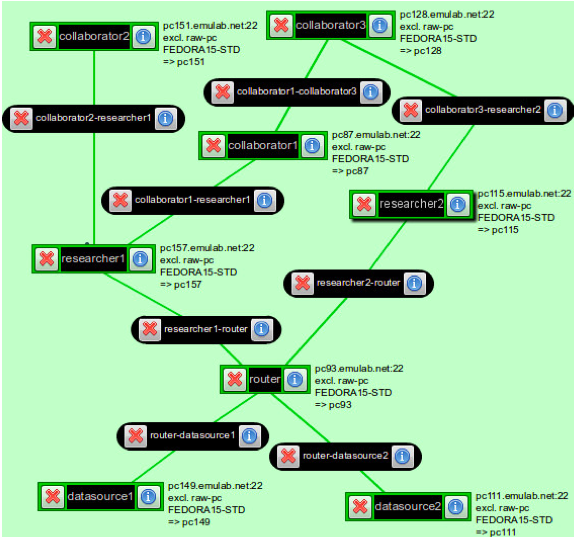
The following is the topology that you will be creating:
- CCNx network for sharing precipitation data:
Prerequisites
- GENI Account:
- Go to Request_GENI_Account for an emulab account and to join a GENI project.
- Or get a GENI Portal account via: Portal Main Site
- SSH
- Unix command line - see the Tools section
- Understanding architectural features of CCNx and other content-centric networking, such as publish-subscribe content distribution, caching at intermediate nodes, and intelligent request forwarding
Tools
- ccnd
The ccnd daemon and its associated software is the reference implementation for the content-centric networking protocol proposed by Project CCNx. This assignment uses release 0.6.1 of the ccnx software package. More information on this package can be found at the Project CCNx web site, http: www.ccnx.org/. The CCN protocol embodied in CCNx is an attempt at a clean-slate architectural redesign of the Internet. - Atmos
The Atmos software package from the NetSec group at Colorado State University will provide named data for your network in the form of precipitation measurements. The Atmos tools are installed in /opt/ccnx-atmos. - NetCDF Tools
NetCDF (Network Common Data Form) is “a set of software libraries and self-describing, machine-independent data formats that support the creation, access, and sharing of array-oriented scientific data.” The NetCDF tools package provides several tools for manipulating NetCDF data, including ncks, a tool for manipulating and creating NetCDF data files; ncdump, a similar tool used for extracting data from NetCDF data files; and ncrcat, a tool for concatenating records from multiple NetCDF data files. - Flack
Flack is a graphical interface for viewing and manipulating GENI resources. It is available on the web at http://www.protogeni.net/trac/protogeni/wiki/Flack. You may use either Omni or Flack to create and manipulate your slices and slivers, but there are certain operations for which you will probably find one or the other easier to use. You will have to use the Flack client to Instrumentize your slices, as you will be directed some exercises.
How to get Help
- If you are using a specific aggregate or tool, you should consider registering in their mailing list. It is a great way to get connected with other GENI users and it is an excellent source of wisdom.
- Send mail to the GENI help list: help@geni.net.
- If you want to chat real-time with other GENI users and ask questions, join us in a GENI chatroom.
Resources
- GENI resources include a variety of computational and network assets, all programmable by experimenters. Resources are made available to experimenters by GENI aggregates. The UnderstandingGENI page has more information in its An Experimenter's View of GENI and GENI Aggregates sections.
Tutorial Instructions
|
|

|
|

|
|
Attachments (1)
- CCNAssignment.png (253.3 KB) - added by 11 years ago.
Download all attachments as: .zip
