| Version 7 (modified by , 11 years ago) (diff) |
|---|
<CCN ASSIGNMENT>

|

|

|
STEPS FOR EXECUTING EXERCISE
Objectives
The objective of this assignment is to familiarize you with named data (sometimes called content-centric) networking. You will specifically learn about:
- The Project CCNx named data networking overlay for IP networks
- Architectural features of CCNx and other content-centric networking, such as publish-subscribe content distribution, caching at intermediate nodes, and intelligent request forwarding
- Using named data networking (NDN) to fetch content from multiple sources
- The effects of cache granularity on bandwidth requirements for content distribution in a named data network
- The usage of intelligent forwarding to reduce bandwidth requirements in a collaborative environment
Tools
- ccnd
The ccnd daemon and its associated software is the reference implementation for the content-centric networking protocol proposed by Project CCNx. This assignment uses release 0.6.1 of the ccnx software package. More information on this package can be found at the Project CCNx web site, http: www.ccnx.org/. The CCN protocol embodied in CCNx is an attempt at a clean-slate architectural redesign of the Internet. - Atmos
The Atmos software package from the NetSec group at Colorado State University will provide named data for your network in the form of precipitation measurements. The Atmos tools are installed in /opt/ccnx-atmos. - NetCDF Tools
NetCDF (Network Common Data Form) is “a set of software libraries and self-describing, machine-independent data formats that support the creation, access, and sharing of array-oriented scientific data.” The NetCDF tools package provides several tools for manipulating NetCDF data, including ncks, a tool for manipulating and creating NetCDF data files; ncdump, a similar tool used for extracting data from NetCDF data files; and ncrcat, a tool for concatenating records from multiple NetCDF data files. - Flack
Flack is a graphical interface for viewing and manipulating GENI resources. It is available on the web at http://www.protogeni.net/trac/protogeni/wiki/Flack. You may use either Omni or Flack to create and manipulate your slices and slivers, but there are certain operations for which you will probably find one or the other easier to use. You will have to use the Flack client to Instrumentize your slices, as you will be directed some exercises.
Exercises
- 3.1 Routing in a Named Data Network
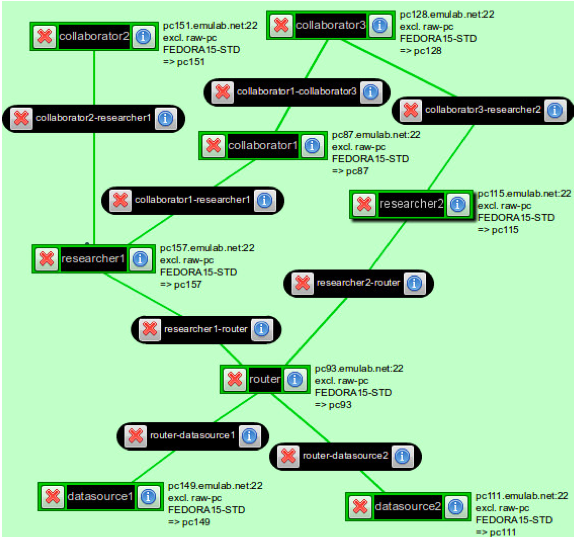
In this exercise, you will build a named data network using the Project CCNx networking daemon ccnd and transfer scientific data in the NetCDF format between data providers and data consumers using routing based on content names. For the purpose of this exercise, the network will use static routes installed at slice setup. Content will be served and consumed by the Atmos software package.
Create a GENI experiment using the precip.rspec file downloaded from here. It will allocate a network such as that depicted in the above figure, with ccnd running on every host and an Atmos server running on the hosts datastore1 and datastore2.
Routing in named data networks takes place using the namespace under which data is stored. In the case of CCNx, the data namespace is a doc/technical/URI.html URI, so it looks similar to the familiar http:// and ftp:// URLs of the World Wide Web. CCNx URIs are of the form ccnx:<path>, where <path> is a typical URI slash-delimited path. The components of the URI path are used by the network to route requests for data from consumers to the providers of that data. Routes have been installed on all of the machines in the network except researcher2, a data consumer node with two connections to the rest of the named data network.- Task 1.1: Finalize Configuration
In this task, you will observe the effects of the missing static routes, and restore them.
Log into the host researcher1. Run the command ccninitkeystore. Issue the command ccndstatus to print the status of the ccnd daemon on the local host. Note the number of content items stored on this host (which should be zero, if you have not yet fetched any content) and the number of interests accepted (under “Interest totals”).
On researcher1, issue a request for precipitation data from 1902/01/01 through 1902/01/02. To do this, use the Atmos tools by invoking:/opt/ccnx-atmos/client.py
Answer the date questions appropriately, and observe that a .nc NetCDF file is created in the current directory containing precipitation data for the specified dates.
Run ccndstatus again, and look at the statistics on the local ccnd. You should see both that a substantial number of content items are cached on this machine (this is the meaning of the “stored” field under “Content items”) and that many more Interests have been seen.
Next, log into researcher2 and issue the same command. Observe that it takes much longer at each step, does not terminate, and no .nc file is created. This is because the ccnd on researcher1 had routes to the precipitation data requested by Atmos, but researcher2 does not. Issue the following command to install the appropriate routes:ccndc add ccnx:/ndn tcp router
Note that the ccndc command takes a content path (ccnx:/ndn) and binds it to a network path (in this case, the host router, which is a valid hostname on the experiment topology, resolving to an IP address on the router machine). It can also be used to delete routes with the del subcommand. - Question 1.1 A:
From researcher2, fetch the same data again (from 1902/01/01 to 1902/01/02), and record the fetch times reported by client.py. It prints out the time take to pull each temporary file along with the concatenation and write time. Then fetch 1902/02/03 to 1902/02/04, and record those fetch times. Fetch 1902/02/03 to 1902/02/04 a second time, and record the new times. Which transfer was longest, and which was shortest? Knowing that each ccnd caches data for a short period of time, can you explain this behavior? - Question 1.1 B:
Browse the content caches and interests seen on various hosts in the network by loading their ccnd status page on TCP port 9695 in your browser (see Section 5, Hints, below). Which hosts have seen interests and have content cached, and why? - Task 1.2: Intent Propagation
This task will demonstrate the behavior of Intent filtering during propagation.
Log into the machine datasource1 and run the command tail -f /tmp/atmos-server.log. Next, run /opt/ccnx-atmos/client.py on both of the hosts researcher1 and researcher2. Enter 1902/01/15 as both start and end date on both hosts, and press <ENTER> on both hosts as close to the same time as possible. - Question 1.2 A:
What intents show up at datasource1 for this request? (Use the timestamps in the server log to correlate requests with specific intents.) Will you observe unique or duplicate intents when both researcher1 and researcher2 fetch the same data at precisely the same time? Why?
- Task 1.3: Explore traffic patterns
For this task, you will start network monitoring services on your slice and observe the data traffic passing between hosts as you fetch various content.
Load your sliver in Flack (if you used your GENI Portal credential to create the topology, then launch Flack from GENI Portal web page) if you have not already done so by refreshing your slices (if necessary) and clicking the slice name in the left pane. Go to the Plugins tab (depicted as a puzzle piece) and click the Instrumentize button. This will take some time, as it adds and configures measurement and monitoring software to your sliver. When it completes, open the INSTOOLS portal in your web browser (Flack will prompt you) and look at the bandwidth usage between the various hosts over time in the INSTOOLS plots.
Log into the host collaborator1 and use the Atmos client to fetch data from 1902/01/28 to 1902/02/03. Look at the INSTOOLS data usage, and observe that data is pulled from both datasource1 and datasource2, through router to researcher1 and ultimately to collaborator1. Record or remember the data rates and volumes from this transaction.
- Task 1.1: Finalize Configuration
In this task, you will observe the effects of the missing static routes, and restore them.
Next: Teardown Experiment
Attachments (2)
- CCNAssignment.png (253.3 KB) - added by 11 years ago.
- topology1.png (115.9 KB) - added by 11 years ago.
Download all attachments as: .zip
