| Version 15 (modified by , 11 years ago) (diff) |
|---|
This page will guide you through Running a Complete Experiment in GENI as seen in the plenary demo!
For this experiment you will need :
- A GENI Account. If you don't have one sign up. If you are at the GEC, get a temporary account.
- An Irods Account
- The tutorial VM. If you do not have it please find us and we will give you a copy.
That's, you are ready to run this experiment!
1. Account Configuration
First we are going to configure all the tools that we will need for this experiment. More specifically we will use :
- Flack to reserve our resources
- GEMINI to instrument our experiment
- OMNI to orchestrate the experiment
We are going to start the configuration in the reverse direction.
1a. Configure Omni on the VM
Omni is already installed in the tutorial VM, but we will need to configure it with your account. Follow these instructions using your personal username, password and passphrase.
Omni scripting for orchestrating the experiment
For orchestrating our experiment we used Omni scripting. We based our script on the readyToLogin.py script. We needed to modify it slightly and then we created a script that uses remote execution to give instructions to the hosts. You can find a copy of our script and the modified ready to login script through these links. For our purpose we placed these two files under /usr/local/bin/gcf/examples/ in the VM.
1b. Configure GEMINI on the VM
These steps are based on these instructions.
- Place your GENI passphrase in a file under
~/.ssl/passwordecho "<passphrase>" > ~/.ssl/password
and verify that it is correctcat ~/.ssl/password
- Add your ssh key to the ssh-agent:
ssh-add ~/.ssh/geni_key
- Add your certificate to firefox. Follow these instructions. Before you start you will need a copy of your cert in
pkc12format. To do that :- login to pgeni.gpolab.bbn.com using your account
- Click the Download certificate link on the left
- Download it in pkc12 format.
1c. Login to Flack
We are going to use Flack a web-based graphical tool for reserving GENI resources. Your first step is to log in to Flack. This video will guide you through the steps of logging in.

|
|
2. Launch your experiment
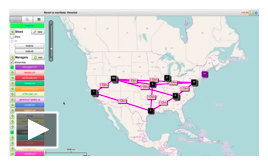
Now that you are logged in to Flack, we are ready to design our experiment. This video will guide you through the process of setting up the resources for the Hello GENI experiment. To complete the setup you will need to save a copy of this file on your computer. This is a Resource Specification (rspec) file that contains a description of this experiment for Flack.
|
|
3. Enable Instrumentation In your Experiment
Now that your slice is ready we will need to instrumentize it. On the VM open a terminal and do:
- Navigate to the right GEMINI folder
cd ~/Tutorials/GEMINI/common
- Run the instrumentize script
instrumentize.py -f ~/.ssl/geni_cert.pem -n <slicename>
This will take a while so be patient until it is done. - Once it is done, use the printed information to login to the [ https://geminiportal.netlab.uky.edu portal].
- Navigate to the graphs. In the map that will appear after you login to the portal
- press on the
GNnode, - select
passive - on the left, navigate to the graphs for
PC1
- press on the
- Run the instrumentize script
4. Run your experiment
You are all set to run your experiment. On the terminal do :
- Go to the location of the script
cd /usr/local/bin/gcf/examples
- Run the script
./race-script.py -a pg-utah <slicename>
The script will first get the information about logging in to your nodes and then it will start executing remote commands to your nodes. It will change the delay of the links and then initiate a UDP and a TCP transfer.- Look at the TCP and UDP graphs as the script is running and notice the performance of UDT and TCP, side by side.
5. Archive your data
Once the script is done, its time to archive the run. Follpw these instructions using your own account information. You can modify the script to test for other values of delay, or bandwidth, or loss rate.
6. Cleanup
Once you are done with your testing, it is time to release the resources. In order to cleanup your slice :
- Press at the Delete button in the bottom of your canvas
- select to delete it at used managers only and confirm your selection.
Wait and after a few moments all the resources will have been released and you will have an empty canvas again. Notice that your slice is still there. There is no way to delete a slice, it will be removed automatically after its expiration date, but remember that a slice is just an empty container so it doesn't take up any resources.
Attachments (3)
- race-scrpt.py (3.8 KB) - added by 11 years ago.
- readyToLogin.py (17.0 KB) - added by 11 years ago.
- race.rspec (7.0 KB) - added by 11 years ago.
Download all attachments as: .zip
