| Version 55 (modified by , 13 years ago) (diff) |
|---|
Table of Contents
Experimentation with GENI
1.0 Why GENI?
GENI might be right for you if your experiment requires:
- More resources than would ordinarily be found in your lab. Since GENI is a suite of infrastructures it can potentially provide you with more resources than is typically found in any one laboratory. This is especially true for compute resources: GENI provides access to large testbeds with hundreds of PCs and to cloud computing resources.
- Non-IP connectivity across resources. Some GENI aggregates allow you to set up Layer 2 connections between resources within the aggregate. Experimenters may install and run their own Layer 3 and above protocols on these resources. It is also possible to setup Layer 2 connections between many GENI aggregates that connect to GENI backbone networks (Internet2 and NLR). You can even set up your network to route through experimenter programmable switches in the GENI backbone.
- A deeply programmable network. GENI has switches in the backbone and at the edges that you can program to set up the network topologies you need and to control flows in your network.
- Geographically distributed resources. Some GENI resources are distributed around the world.
- Reproducibility. You can get exclusive access to certain GENI resources including CPU resources and network resources. This gives you control over your experiment's environment and hence the ability for you and others to repeat experiments under identical or very similar conditions.
2.0 An Experimenter's View of GENI
GENI is a suite of infrastructures for networking and distributed systems experimentation. GENI supports at-scale experimentation on shared, heterogeneous, highly instrumented infrastructure and enables deep programmability throughout the network.
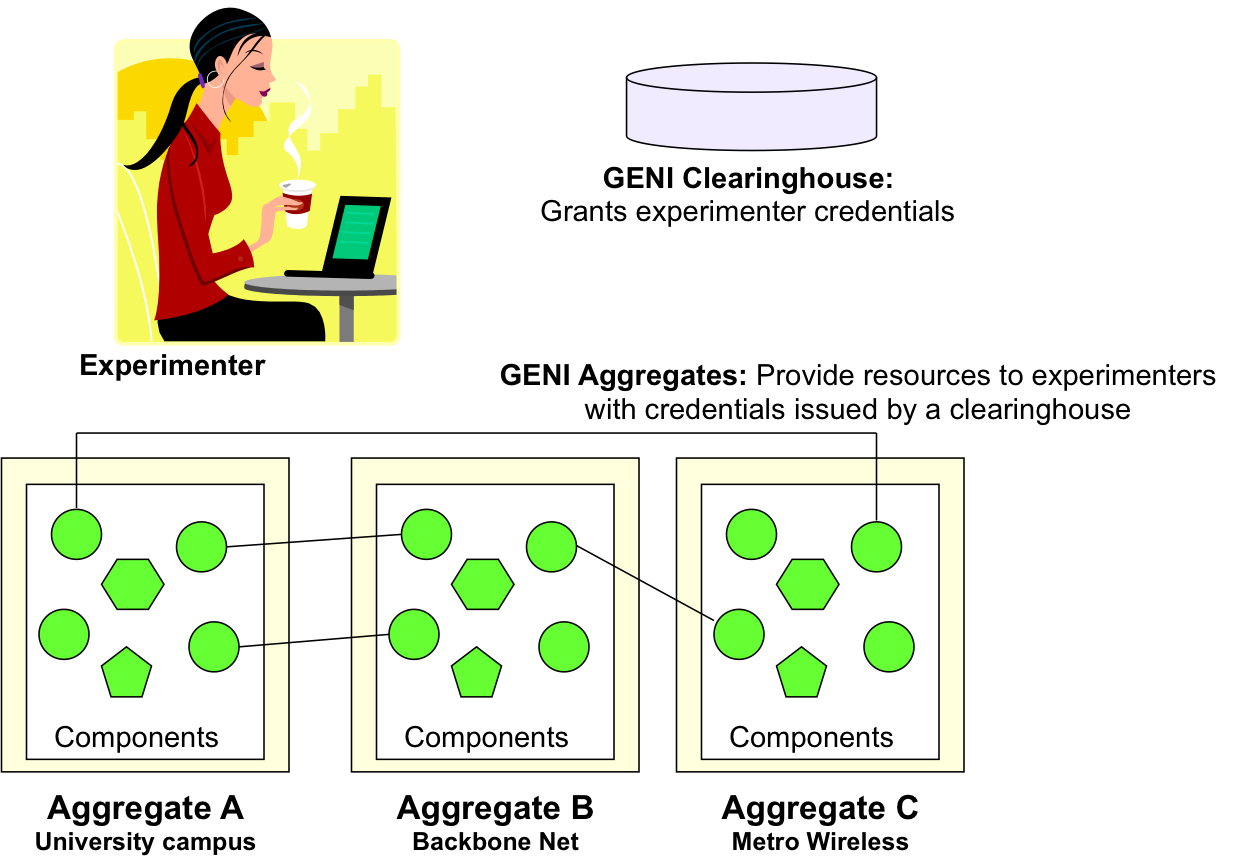
As an experimenter you will need to know about GENI clearinghouses and GENI aggregates. A GENI clearinghouse authenticates experimenters and issues them credentials needed to obtain GENI resources for experimentation.
GENI aggregates provide resources to experimenters with GENI credentials. GENI has a number of different aggregates that provide a variety of resources for experimentation. An important aspect of planning your experiment is deciding what resources you need (type and numbers) and which aggregates might meet your needs.
The following figure illustrates the role of GENI clearinghouses and aggregates:
You will also need to know about GENI slices. A slice holds a collection of computing and communications resources capable of running an experiment or a wide area service. An experiment is a researcher-defined use of resources in a slice; an experiment runs in a slice. A researcher may run multiple experiments using resources in a slice, concurrently or over time.
3.0 GENI Aggregates
The following table lists GENI aggregates that are currently available for use by experimenters and the networks (GENI backbone network or the Internet) to which they connect. GENI has two backbone networks: Internet2 and National Lambda Rail (NLR). The Internet2 backbone provides 1Gbps of dedicated bandwidth for GENI experiments and the NLR backbone provides up to 30Gbps of non-dedicated bandwidth. Some aggregates that connect to GENI backbone networks may be connected to other resources on the network using Layer 2 VLANS, giving experimenters the option of running non-IP based Layer 3 and above protocols.
3.1 GENI aggregates currently available to experimenters:
GENI Aggregate Providers: Please report errors and omissions in the table below to Vic Thomas
| Aggregate | Description | Compute Resources | Programmable Network | Wireless | Network Connectivity | Experimenter Tools |
|---|---|---|---|---|---|---|
| PlanetLab | Testbed consisting of 1090 nodes at 513 sites around the world | Virtual machines on PlanetLab nodes | No | No | Internet | Gush, Omni, Raven, SFI |
| GPO Lab myPLC | PlanetLab installation consisting of 5 multi-homed nodes | Virtual machines on PlanetLab nodes | No | No | Internet2: IP; NLR: IP; Internet | Gush, Omni, SFI |
| Utah ProtoGENI | Over 500 co-located PCs that can be loaded with an experimenter specified OS image and connected in arbitrary topologies | Complete PCs or virtual machines on PCs | PCs can be set up as routers, plus experimenter-controllable switches (HP ProCurves) | 60 nodes with 2 WiFi cards each, plus software-defined radio peripherals (USRP2) | Internet2: IP and Layer 2; Internet | ProtoGENI Tools, Gush |
| Kentucky ProtoGENI | Over 50 co-located PCs that can be loaded with an experimenter specified OS image and connected in arbitrary topologies. Strong instrumentation capabilities | Complete PCs or virtual machines on PCs | PCs can be set up as routers | No | Internet2: IP and Layer 2; Internet | ProtoGENI Tools, Instrumentation Tools |
| GPO Lab ProtoGENI | 11 co-located PCs that can be loaded with an experimenter specified OS image and connected in arbitrary topologies | Complete PCs | PCs can be set up as routers | No | Internet2: IP and Layer 2; NLR: IP and Layer 2; Internet | ProtoGENI Tools, Gush |
| Deter |
GENI Aggregate Providers: Please report errors and omissions in the table above to Vic Thomas
3.2 GENI aggregates that will be available soon, with links to their GENI project page:
GpENI Network testbed centered on a Midwest US regional optical network between The University of Kansas, Kansas State University, University of Nebraska – Lincoln, and University of Missouri – Kansas City, supported with optical switches from Ciena interconnected by Qwest fiber infrastructure. GENICloud Brings OpenCirrus and Ecalyptus-based cloud computing resources to GENI experimenters. PrimoGENI Integrate a large-scale, real-time network simulator (PRIME) into ProtoGENI, enabling slices involving both physical and simulated networked components. CMU Homenet Nodes Nodes on non-controlled networks placed in residences such as apartments. MAX Regional optical network consisting of wavelength-selectable switches, 10Gbps Ethernet switches, PlanetLab nodes, OpenFlow switches, NetFPGA hosts, and connections to ProtoGENI, Internet2 ION, NLR. MAX Aggregate CMU Wireless Emulator A wireless network emulator that accurately emulates wireless signal propagation in a physical space. ORCA/BEN Network consisting of several segments of dark fiber and includes a reconfigurable fiber switch (layer 0) to generate different physical topologies, out of band network management to access equipment at PoPs and remote power management for resetting and powering down of experimental equipment. Data Intensive Cloud Cloud-based environment for data-intensive experiments from start (the data collection point) to finish (processing and archiving). Rutgers WiMAX GENI-enabled WiMAX base station at Rutgers University. NY Poly WiMAX GENI-enabled WiMAX base station at Polytechnic Institute of New York University. UCLA WiMAX GENI-enabled WiMAX base station at the University of California, Los Angeles to support the deployment and testing of vehicular services and applications in the Campus Vehicular Testbed (C-VeT). GPO Lab WiMAX GENI-enabled WiMAX base station at the BBN Campus, Cambridge, MA.
3.3 Picking Resources for Your Experiment
As you plan your experiment you will want to consider:
- The degree of control you need over your experiment. Do you need to tightly control the resources (CPU, bandwidth, etc.) allocated to your experiment or will best-effort suffice. If you need a tightly controlled environment you might want to consider the U. of Utah ProtoGENI aggregate that allocate entire PCs that can be connected in arbitrary topologies.
- The desired network topology. Does your experiment have to be geographically distributed? What kinds of connectivity do you need between these geographically distributed locations. Almost all aggregates can connect using IP connectivity over the Internet. Many aggregates connect to one of the GENI backbones and allow you to set up IP connections with other resources on the backbone. This will give you a bit more control over the network. Some aggregates provide Layer 2 connectivity over a GENI backbone i.e. you can set up vlans between these aggregates and other resources on the backbone network. This allows you to run non-IP protocols across between the aggregate and other resources.
- The desired control over network flows. If you need to manage network traffic to/from an aggregate you might want to use aggregates that connect to a GENI backbone using OpenFlow switches or set up vlans to these aggregates through the ProtoGENI Backbone Nodes or the SPP Nodes.
- The number of resources you need from an aggregate. Aggregates vary from small installations such as the GPO Lab ProtoGENI aggregate that consists of eleven nodes to the PlanetLab and ProtoGENI aggregates that consist of hundreds of nodes.
The GENI Project Office is happy to help find the best match of resources for your experiments. Please contact help@geni.net for assistance.
4.0 Experimenter Tools
4.1 Experiment Control Tools
GENI experiment control tools are used to create slices, add or remove resources to slices, and delete slices. Some tools may also help with the installation of experimenter specified software into resources in slices; starting, pausing, resuming and stopping the execution of an experiment; and monitoring of the resources in slices for failures. Examples of GENI experiment control tools include Gush, Omni, PlanetLab SFI and ProtoGENI Tools.
In addition to these experiment control tools, individual aggregates provide experimenters with additional tools to install and manage software on their resources. For example, the Million Node GENI aggregate provides a set of tools to manage the virtual machines it proves as computing resources.
4.2 Instrumentation and Measurement Tools
GENI instrumentation tools are currently aggregate specific. Examples of such tools include Instrumentation Tools for the Kentucky ProtoGENI aggregate, Owl for the PlanetLab aggregate and OMF/OML for the ORBIT aggregate.
5.0 Getting Access to GENI
To use GENI for experimentation please contact help@geni.net.
6.0 Tutorials
For a tutorial on using Omni tools to run experiments on GENI, see http://groups.geni.net/geni/wiki/GENIExperimenter.
For a tutorial on using ProtoGENI Tools to run experiments on GENI, see http://www.protogeni.net/trac/protogeni/wiki/Tutorial.
Attachments (4)
-
GENIComponentsPicture-2.png (131.7 KB) - added by 13 years ago.
Clearinghouses and Aggregates
-
ResourceTable.png (201.4 KB) - added by 13 years ago.
Table of aggregates providing compute resources
-
Internet2-Map.jpg (449.5 KB) - added by 13 years ago.
Internet2 Toplogy (from http://www.internet2.edu/network/)
-
NLR-Map.jpg (166.2 KB) - added by 13 years ago.
NLR Topology (from http://noc.nlr.net/nlr/maps_documentation/nlr_wavenet_documentation.html)
Download all attachments as: .zip