| Version 6 (modified by , 11 years ago) (diff) |
|---|
GIMI Init Script in Exper Mgmt Envir (EME)
1) Goals
The GIMI init script in EME aims to interactively initialize the experiment environment for each user-running experiment.
2) Tasks for Init Script
- Get experiment name from user, and derive experiment-id
- Retrieve project_id from omni or CH
- Get list of slice_ids from omni or CH, select slice_id for this experiment
- Get sliver_manifests_rspecs from AMs
- Parse sliver_manifest_rspec to get slice_node_names (fully qualified)
- Select default GIMI Portal, or override
- Setup GSAS (iRODS) structure
- derive defaults for user_irods_home_directory and
- target(exper)-directory, or override
- derive and load proj_exper_step_descriptors
- load template OMF scripts
- get user_irods_target(exper)_iticket, assigned to selcted GIMI Portal agent
- Push sliver_manifest_rspecs and exp_descriptors to iRODS
- Holding user_credentials, config GIMI Portal exper info:
- user_identity
- experiment_id,
- project_id, and slice_id
- slice_node_names
- sliver_manifest_rspecs
- user_irods_home_directory
- user_irods_target(exper)_directory
- user_irods_target(exper)_iticket
3) GIMI Init installation
This session describes the installation procedure The script can be obtained from github: https://github.com/geni-gimi/GIMI/tree/master/gimiInit
4) How to run
This session describes the running and consequences of gimi_init
5) Experiment descriptor and iRODS
This script sets up an experiment collection within the user's iRODS account. The user and GIMI will store objects associated with this experiment in this collection. In the example below the experiment collection is called “myExperiment” .
Template OMF scripts are also added to the user's iRODS account in their own folder titled “experimentTemplates” the first time a user runs this script.
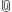
Descriptor files are generated which collect metadata about the experiment and objects being pushed into iRODS. These descriptor files are used to help organize and search for data. "project.xml" and “experiment.xml” are descriptor files created in each experiment collection.
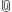
The user is asked if he would like the manifest rspecs from his slice to be added to this experiment collection in iRODS. If the user says yes, a new collection is created within the experiment collection. The manifest rspecs are then pushed to this new collection and descriptor files are created. In this example "TwoVMs" and "ThreeVMs" are manifest rspecs and "artifact.xml" and “step.xml” are descriptor files.
Attachments (4)
- MainCollection.png (62.5 KB) - added by 11 years ago.
- ExperimentCollection.png (70.1 KB) - added by 11 years ago.
- ManifestCollection.png (75.9 KB) - added by 11 years ago.
- ExperimentTemplates.png (94.3 KB) - added by 11 years ago.
Download all attachments as: .zip
