| Version 11 (modified by , 11 years ago) (diff) |
|---|
GIMI Init Script in Exper Mgmt Envir (EME)
1) Goals
The GIMI init script in EME aims to interactively initialize the experiment environment for each user-running experiment.
2) Tasks for Init Script
- Get experiment name from user, and derive experiment-id
- Retrieve project_id from omni or CH
- Get list of slice_ids from omni or CH, select slice_id for this experiment
- Get sliver_manifests_rspecs from AMs
- Parse sliver_manifest_rspec to get slice_node_names (fully qualified)
- Select default GIMI Portal, or override
- Setup GSAS (iRODS) structure
- derive defaults for user_irods_home_directory and
- target(exper)-directory, or override
- derive and load proj_exper_step_descriptors
- load template OMF scripts
- get user_irods_target(exper)_iticket, assigned to selcted GIMI Portal agent
- Push sliver_manifest_rspecs and exp_descriptors to iRODS
- Holding user_credentials, config GIMI Portal exper info:
- user_identity
- experiment_id,
- project_id, and slice_id
- slice_node_names
- sliver_manifest_rspecs
- user_irods_home_directory
- user_irods_target(exper)_directory
- user_irods_target(exper)_iticket
3) GIMI Init installation
This section describes the installation procedure The script can be obtained from github: https://github.com/geni-gimi/GIMI/tree/master/gimiInit
4) How to run
Open a terminal window and run:
python gimi_init.py
The output should look like this:
Welcome to the GIMI initialization script, please make sure to install omni and set up your credential before running this script Are you creating a new experiment?
If you are creating a new experiment enter “yes”.
If you are updating an existing experiment enter “no” and continue to these instructions link.
If you entered “yes” your, output should be the following:
Enter your preferred experiment path (please use absolute path, e.g., /home/geni/your/dir):
You will now need to input the path to the directory on your VM that you would like to store your experiment information in.
For example you might want to store it your experiment files here:
/home/geni/GIMIexperiment
If the path doesn't already exist your output will look like this:
Path doesn't exist, do you want me to create directory for you? (Yes or No)
Enter “yes” if you would like to create that directory. The init script will now ask you a few question about yourself to be put in the descriptor files that will accompany your experiment files.
Tell us about yourself: enter your name:
It will now ask for the organization you are a part of. This could be a company or university.
Your organization:
Next iRODS will be initialized. You should already have created the file “~/.irods/.irodsEnv”. If you have not done this you will need to answer the iRODS questions with information from your iRODS account which you created in the GENI Portal. You can get additional information at http://groups.geni.net/geni/wiki/HowToUseiRODS HowToUseiRODS.
Your next output will list your slices and ask you to choose which slice you are using for this experiment.
Your user name is ... Your active slice names: ... ... Please enter your preferred slice name for this experiment:
After you enter which slice you would like to use, the output will tell you which slice you chose and which project that slice is a member of. Then it will ask you to name your experiment.
Your slice name is: ... Your project ID is: … Enter your experiment name:
You will then be told the experiment ID. Then the output will look like this:
Retrieving the manifest rspec... Checking resources from aggregates... (this might take some time)
Depending on which aggregates your slice has resources, you should get additional output about where your resources were found. Here is an example:
Found resource in aggregate: https://boss.instageni.gpolab.bbn.com:12369/protogeni/xmlrpc/am/2.0
Next the OMF template scripts will be pushed to iRODS if they are not already there & iRODS will be set up for the experiment. You will also be given a ticket for the experiment directory that can be used to give other users access.
iRODS initial collection setup was successful Ticket for new directory: ... Ticket expiration date is set to: ... Do you want to push manifest to iRODS? (Yes or No)
You should push manifest to iRODS if you have already reserved your resources so enter “yes”. You should get the following output:
Manifest has been pushed to iRODS Pushing experiment information to GIMI Experiment Registry... Saving JSON to file: …
This will be followed by the JSON files that are pushed to the registery. At the end of all this output you should see:
Pushed data to registry successfully
You have successfully run the GIMI initialization script.
5) Experiment descriptor and iRODS
Initial Experiment Collection
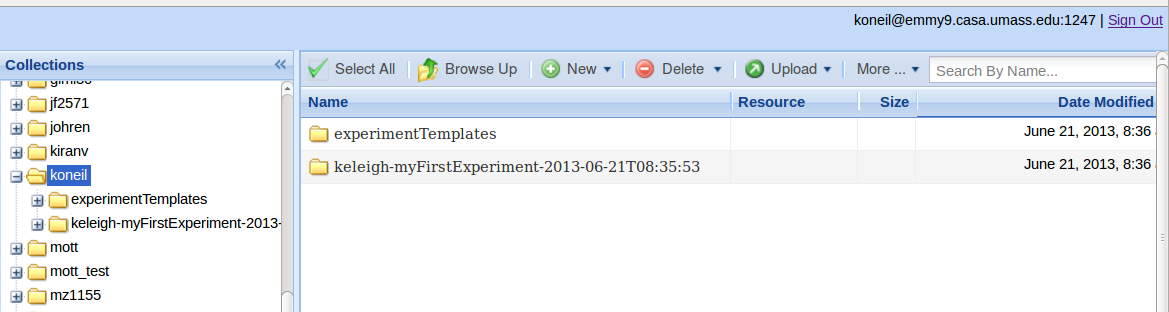
This script sets up an experiment collection within the user's iRODS account. The user and GIMI will store objects associated with this experiment in this collection. In the example below the experiment collection is called “keleigh-myFirstExperiment-2013-06-21T08:35:53” .
Descriptor Files
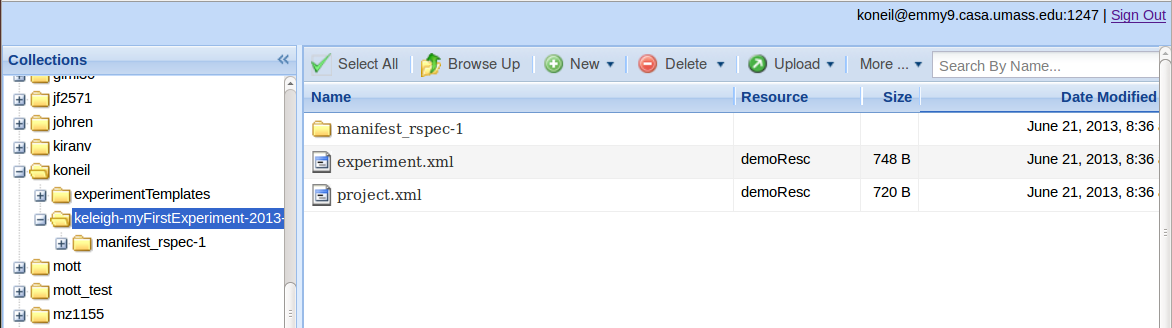
Descriptor files are generated which collect metadata about the experiment and objects being pushed into iRODS. These descriptor files are used to help organize and search for data. "project.xml" and “experiment.xml” are descriptor files created in each experiment collection.
Manifest Rspecs
The user is asked if he would like the manifest rspecs from his slice to be added to this experiment collection in iRODS. If the user says yes, a new collection is created within the experiment collection. The manifest rspecs are then pushed to this new collection and descriptor files are created. In this example "manifest-keleigh-myFirstExperiment-2013-06-21T08:35:53.rspec" is a manifest rspec and "artifact.xml" and “step.xml” are descriptor files.
Experiment Templates
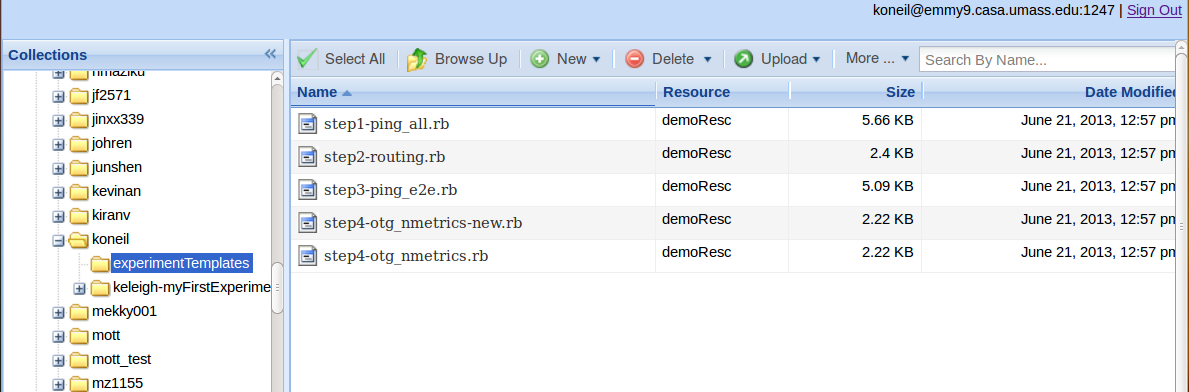
Template OMF scripts are added to the user's iRODS account in their own folder titled “experimentTemplates” the first time a user runs this script. These OMF scripts are also available in Labwiki
Attachments (4)
- MainCollection.png (62.5 KB) - added by 11 years ago.
- ExperimentCollection.png (70.1 KB) - added by 11 years ago.
- ManifestCollection.png (75.9 KB) - added by 11 years ago.
- ExperimentTemplates.png (94.3 KB) - added by 11 years ago.
Download all attachments as: .zip