INSTOOLS Project Status Report
Period: Post-GEC11 Report
I. Major accomplishments
The following highlights our accomplishments during the last reporting period.
A. Milestones achieved
During the last reporting period we achieved the following milestones:
- (milestone s3.d) We identified multiple GENI experiments that will use our Instrumentation Tools. We made contact with these experimenters and have been providing support as they begin to use the tools. We established a web site with documentation/tutorials/examples to help experimenters get started.
- (milestone s3.e) We gave a tutorial of the instrumentation tools (together with the Utah ProtoGENI tutorial) at GEC 11 to experimenters and educators.
- (milestone s3.f) We made the latest stable version of our software available through the ProtoGENI GIT repository.
- Other achievements include:
- Modifying our design to separate out the user interface from the backend services that deploy the instrumentation infrastructure, thereby enabling a wider range of user interfaces.
- Integrating the INSTOOLS software into the FLACK client/GUI.
- Adding several enhancements to our GENI Monitoring Portal (GMP).
- Simplifying the authentication process for users.
- Adding support for creating archives on the UK Archive Service (UKAS) and the CNRI archive service.
- Adding support for both version 1 and version 2 RSPECs.
- Adding support for Ubuntu Node OS images.
- Improving the robustness/resilience of the tools to errors.
B. Deliverables made
- We made the latest stable version of our software available via the ProtoGENI GIT repository.
- We posted new documentation about INSTOOLS at http://groups.geni.net/geni/wiki/INSTOOLSSummary
- We posted a tutorial providing step-by-step instructions on how to use INSTOOLS at http://www.netlab.uky.edu/p/instools/
II. Description of work performed during last quarter
The following provides a description of our activities and results during the last report period.
A. Activities and findings
During the last reporting period we made a major effort to integrate the instrumentation tools with the ProtoGENI FLACK client.
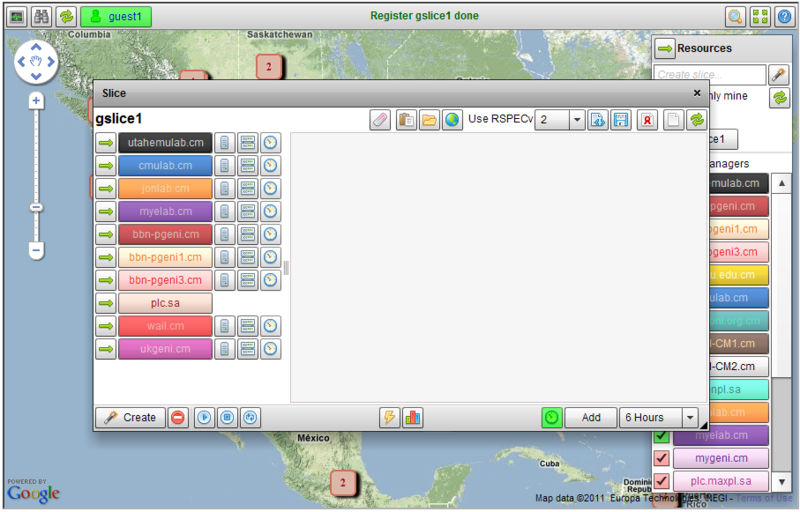
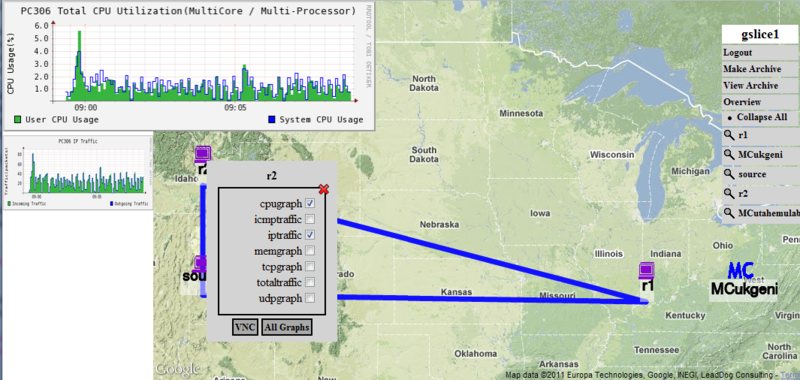
Instead of requiring users to run a set of scripts to instrumentize their experiment, we modified the FLACK GUI so that users can instrumentize their slice simply by clicking on a button in the GUI. In particular, we added two new buttons to the FLACK GUI (Figure 1), one to add instrumentation to a slice (Figure 2) and the other to access the portal site (Figure 3). To instrumentize an existing slice, a user simply needs to click on the instrumentize button in the FLACK GUI. The GUI will then talk with the backend instrumentation servers to instrumentize the slice. While this is occuring, information about the progress/status of the instrumentize process is sent back to the GUI and can be viewed by the user. After the slice has been instrumentized, the Go-to-portal button (Figure 3) is enabled, allowing the user to view the topology and the traffic moving across various nodes and links. The user can pick any node or link in the experiment to observe the measurement data he/she is interested in, as shown in Figure 4.
Figure 2: The Button for Instrumentation.
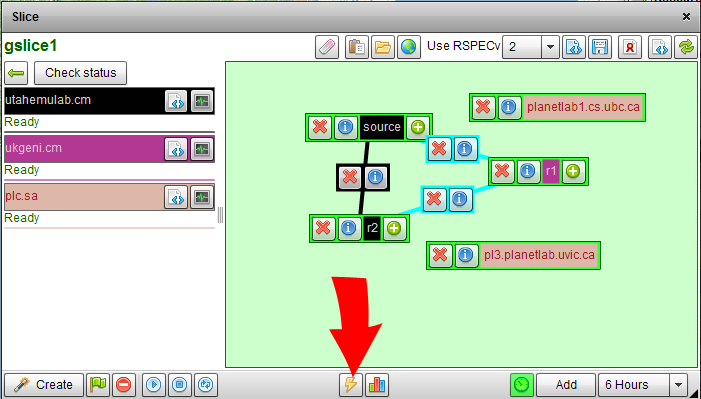
Figure 3: The Button for the Portal.

The integration of the INTOOLS with the FLACK interface greatly simplifies the instrumentation process for users, making it much easier to use the instrumentation tools. From a user's perspective, the integration also enables a sort of single sign-on service in which the user authenticates to the FLACK client, but then the FLACK client and the backend instrumentation manager handles all the authentication on behalf of the user to the long lists of services that comprise the instrumentation system. The portal button on the FLACK client takes the user directly to our GENI Monitoring Portal (GMP) so that the user does not need to remember URLs or have to login to the GMP on their own.
To integrate the instrumentation tools with the FLACK client, we redesigned our software so that the installation and deployment of the instrumentation software is done by a backend instrumentation manager that, like a component manager, is responsible for allocating and setting up instrumentation infrastructure. As a result, the user interface components of our system could be separated out and integrated into the FLACK client. This involved rewriting our Python scripts as flash code making ProtoGENI API calls as well as XML-RPC calls to our instrumentation manager (IM). We currently run an instrumentation manager (IM) at each aggregate in concert with the component manager at each aggregate. The IM is responsible for sshing to experimental nodes, running initialization scripts, setting up the MC nodes, etc and, like a component manager, has the authority needed to carry out these tasks.
The integration of INSTOOLS with the FLACK client is one of our efforts to make the INSTOOLS interface more user friendly and attract more experimenters to use our tools. During this period of time, with the help of GPO, we identified and provided support for experiments (e.g, RIT, Buffalo) to use our instrumentation tools. We updated our web site to reflect the current status of the project and provided documentation/tutorials/examples to help experimenters use our tools better.
On another front, we also connected our INSTOOLS system with two archival systems: (1) the University of Kentucky Archive Service (UKAS) and (2) the CNRI Archive Service. (We have begun discussions with the GMOC to connect to their archive service, but do not yet have support for that in our implementation).
Our UKAS is the most powerful of the services because it not only provides a repository for data, but it also provides a computational environment where we can recreate the same look-and-feel the user had when viewing the live data. In particular, we use OpenVZ containers to provide a virtualized environment in which to run the drupal content management systems that was running on the MC at the the time the archive was taken. As a result, the user can visit the same web pages that were offered by the MC at the time the archive was taken.
We have also incorporated support for the CNRI archive service and its concept of workspaces. In particular, our system is able to store the data files containing measurements information into a CNRI workspace associated with a slice. After the data has been stored in the workspace, the system adds the necessary metadata needed by the CNRI archive to move the measurement files from the workspace to the CNRI archive for permanent storage. The files can then be access via the CNRI web interface.
Together with our Utah colleagues, we gave a tutorial on the instrumentation tools at GEC 11 to experimenters and potential users of our tools. The response to the integrated tool was extremely positive, and we have already begun to see several user using the tool since the GEC.
Finally, as in past reporting periods, we upgraded our ProtGENI software to incorporate the latest changes made by the Utah group, making the necessary changes to our INSTOOLs software in order to run on the upgraded aggregate. We also added support for both version 1 and version 2 RSPECs, we added support for Ubuntu Node OS images (something requested by many users), and we performed quite a bit of testing in preparation for the tutorial that greatly improved the robustness/resilience of our tools to errors. The latest stable version of our software can be obtained via the ProtoGENI GIT repository.
B. Project participants
The following individuals have helped with the project in one way or another during the last quarter:
- Jim Griffioen - Project PI
- Zongming Fei - Project Co-PI
- Hussamuddin Nasir - Technician/programmer
- Xiongqi Wu - Research Assistant
- Jeremy Reed - Research Assistant
- Lowell Pike - Network administrator
- Woody Marvel - Assists in Emulab administration
C. Publications (individual and organizational)
None this quarter.
D. Outreach activities
- We gave a tutorial about INSTOOLS at GEC 11.
- We have been providing support for the early adopters of our tools.
E. Collaborations
Most of our collaborations continue to be with the Utah ProtoGENI team, and we continue to be actively involved in the bi-weekly meetings of the ProtoGENI cluster.
F. Other Contributions
Attachments (4)
-
flack.png (250.3 KB) - added by 13 years ago.
Flack Client
-
goto_instrument.png (58.3 KB) - added by 13 years ago.
Instrumentize Button
-
goto_portal.png (61.9 KB) - added by 13 years ago.
Portal Button
-
portal.png (415.6 KB) - added by 13 years ago.
GENI Monitoring Portal (GMP)
Download all attachments as: .zip